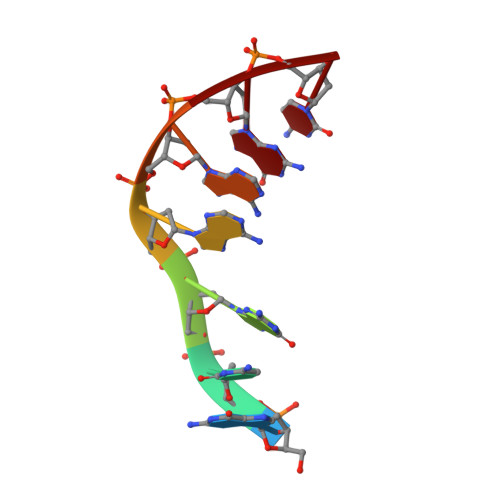
Structures of d(GCGAAGC) and d(GCGAAAGC) (tetragonal form): a switching of partners of the sheared G.A pairs to form a functional G.AxA.G crossing.
Sunami, T., Kondo, J., Hirao, I., Watanabe, K., Miura, K., Takenaka, A.(2004) Acta Crystallogr D Biol Crystallogr 60: 422-431
- PubMed: 14993665 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444903028415
- Primary Citation Related Structures:
1UB8, 1UE4 - PubMed Abstract:
The DNA fragments d(GCGAAGC) and d(GCGAAAGC) are known to exhibit several extraordinary properties. Their crystal structures have been determined at 1.6 and 1.65 A resolution, respectively. Two heptamers aligned in an antiparallel fashion associate to form a duplex having molecular twofold symmetry. In the crystallographic asymmetric unit, there are three structurally identical duplexes. At both ends of each duplex, two Watson-Crick G.C pairs constitute the stem regions. In the central part, two sheared G.A pairs are crossed and stacked on each other, so that the stacked two guanine bases of the G.AxA.G crossing expose their Watson-Crick and major-groove sites into solvent, suggesting a functional role. The adenine moieties of the A(5) residues are inside the duplex, wedged between the A(4) and G(6) residues, but there are no partners for interactions. To close the open space on the counter strand, the duplex is strongly bent. In the asymmetric unit of the d(GCGAAAGC) crystal (tetragonal form), there is only one octamer chain. However, the two chains related by the crystallographic twofold symmetry associate to form an antiparallel duplex, similar to the base-intercalated duplex found in the hexagonal crystal form of the octamer. It is interesting to note that the significant difference between the present bulge-in structure of d(GCGAAGC) and the base-intercalated duplex of d(GCGAAAGC) can be ascribed to a switching of partners of the sheared G.A pairs.
- Graduate School of Bioscience and Biotechnology, Tokyo Institute of Technology, Yokohama 226-8501, Japan.
Organizational Affiliation: