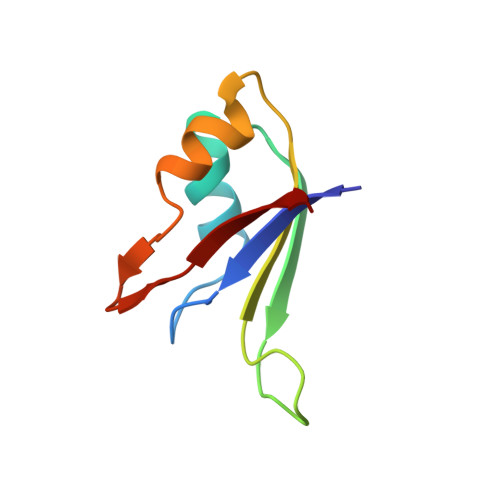
Origin of higher affinity to RNA of the N-terminal RNA-binding domain than that of the C-terminal one of a mouse neural protein, musashi1, as revealed by comparison of their structures, modes of interaction, surface electrostatic potentials, and backbone dynamics
Miyanoiri, Y., Kobayashi, H., Imai, T., Watanabe, M., Nagata, T., Uesugi, S., Okano, H., Katahira, M.(2003) J Biological Chem 278: 41309-41315
- PubMed: 12907678 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M306210200
- Primary Citation Related Structures:
1UAW - PubMed Abstract:
Musashi1 is an RNA-binding protein abundantly expressed in the developing mouse central nervous system. Its restricted expression in neural precursor cells suggests that it is involved in maintenance of the character of progenitor cells. Musashi1 contains two ribonucleoprotein-type RNA-binding domains (RBDs), RBD1 and RBD2, the affinity to RNA of RBD1 being much higher than that of RBD2. We previously reported the structure and mode of interaction with RNA of RBD2. Here, we have determined the structure and mode of interaction with RNA of RBD1. We have also analyzed the surface electrostatic potential and backbone dynamics of both RBDs. The two RBDs exhibit the same ribo-nucleoprotein-type fold and commonly make contact with RNA on the beta-sheet side. On the other hand, there is a remarkable difference in surface electrostatic potential, the beta-sheet of RBD1 being positively charged, which is favorable for binding negatively charged RNA, but that of RBD2 being almost neutral. There is also a difference in backbone dynamics, the central portion of the beta-sheet of RBD1 being flexible, but that of RBD2 not being flexible. The flexibility of RBD1 may be utilized in the recognition process to facilitate an induced fit. Thus, comparative studies have revealed the origin of the higher affinity of RBD1 than that of RBD2 and indicated that the affinity of an RBD to RNA is not governed by its fold alone but is also determined by its surface electrostatic potential and/or backbone dynamics. The biological role of RBD2 with lower affinity is also discussed.
- Department of Environment and Natural Sciences, Graduate School of Environment and Information Sciences, Yokohama National University, 79-7 Tokiwadai, Hodogaya-ku, Yokohama 240-8501, Japan.
Organizational Affiliation: