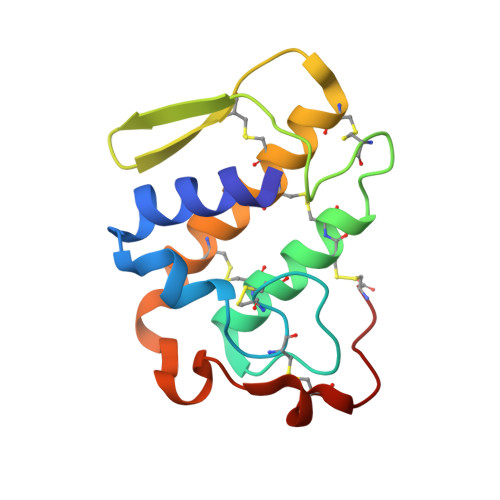
Crystal structure of an acidic platelet aggregation inhibitor and hypotensive phospholipase A(2) in the monomeric and dimeric states: insights into its oligomeric state
Magro, A.J., Murakami, M.T., Marcussi, S., Soares, A.M., Arni, R.K., Fontes, M.R.(2004) Biochem Biophys Res Commun 323: 24-31
- PubMed: 15351695 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2004.08.046
- Primary Citation Related Structures:
1U73, 1UMV - PubMed Abstract:
Phospholipases A2 belong to the superfamily of proteins which hydrolyzes the sn-2 acyl groups of membrane phospholipids to release arachidonic acid and lysophospholipids. An acidic phospholipase A2 isolated from Bothrops jararacussu snake venom presents a high catalytic, platelet aggregation inhibition and hypotensive activities. This protein was crystallized in two oligomeric states: monomeric and dimeric. The crystal structures were solved at 1.79 and 1.90 angstroms resolution, respectively, for the two states. It was identified a Na+ ion at the center of Ca2+-binding site of the monomeric form. A novel dimeric conformation with the active sites exposed to the solvent was observed. Conformational states of the molecule may be due to the physicochemical conditions used in the crystallization experiments. We suggest dimeric state is one found in vivo.
- Departamento de Física e Biofísica, Instituto de Biociências, UNESP, Botucatu-SP, Brazil.
Organizational Affiliation: