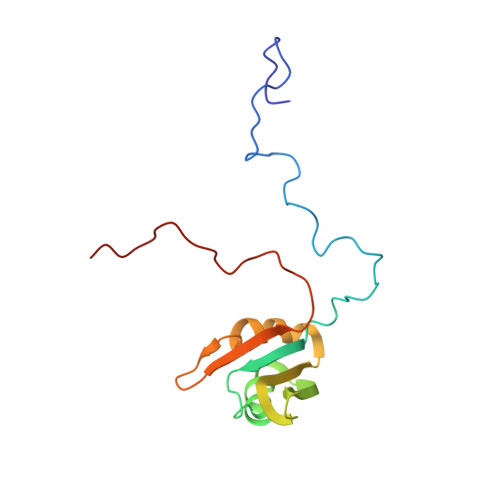
NMR Structural Study of TcUBP1, a Single RRM Domain Protein from Trypanosoma cruzi: Contribution of a beta Hairpin to RNA Binding
Volpon, L., D'Orso, I., Young, C.R., Frasch, A., Gehring, K.(2005) Biochemistry 44: 3708-3717
- PubMed: 15751947 Search on PubMed
- DOI: https://doi.org/10.1021/bi047450e
- Primary Citation Related Structures:
1U6F - PubMed Abstract:
TcUBP1 is a trypanosome cytoplasmic RNA-binding protein containing a single, conserved RNA-recognition motif (RRM) domain involved in selective destabilization of U-rich mRNAs such as the Trypanosoma cruzi small mucin gene family mRNA, TcSMUG. TcUBP1 binds specific transcripts in vivo and co-localizes in the perinuclear part of the cell with components of the mRNA-stability determinant pathway such as poly(A)-binding protein 1 (PABP1) and TcUBP2, a closely related RRM-containing protein. In TcUBP proteins, the RRM domain is flanked by N-terminal Gln-rich and C-terminal Gly-Gln-rich extensions, which are involved in protein-protein interactions. In this work, we determined the solution structure of the TcUBP1 RRM domain by nuclear magnetic resonance (NMR) spectroscopy. The domain has a characteristic betaalphabetabetaalphabeta fold, consisting of a beta sheet composed of four antiparallel betastrands and two alpha helices packed against one face of the beta sheet. A unique aspect of TcUBP1 is the participation of a beta hairpin (beta4-beta5) in the beta sheet, resulting in an enlarged RNA-binding surface. Detailed analysis of the TcUBP1 interaction with a short single-stranded RNA derived from the 3' UTR of TcSMUG was carried out by titration experiments using both NMR spectroscopy and isothermal titration calorimetry. This analysis revealed that amino acids located within the beta hairpin (beta4-beta5) contribute to complex formation. This enlarged protein-RNA interface could compensate for the lack of additional RNA-binding domains in TcUBP1, as observed in many other RRM-containing proteins. The structure of TcUBP1 reveals new aspects of single RRM-RNA interactions and insight into how N- and C-terminal extensions can contribute to RNA binding.
- Department of Biochemistry, McGill University, McIntyre Medical Science Building, 3655 Promenade Sir-William-Osler, Montreal, Québec, H3G 1Y6, Canada.
Organizational Affiliation: