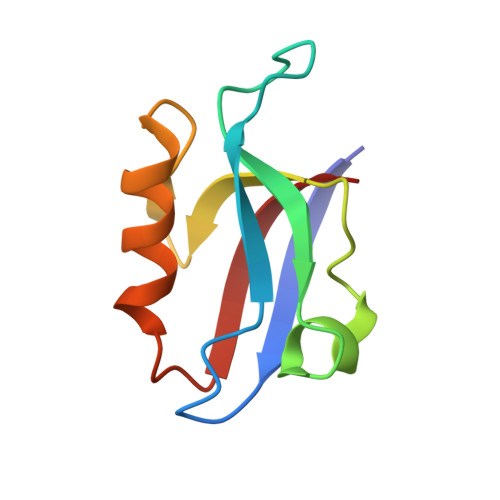
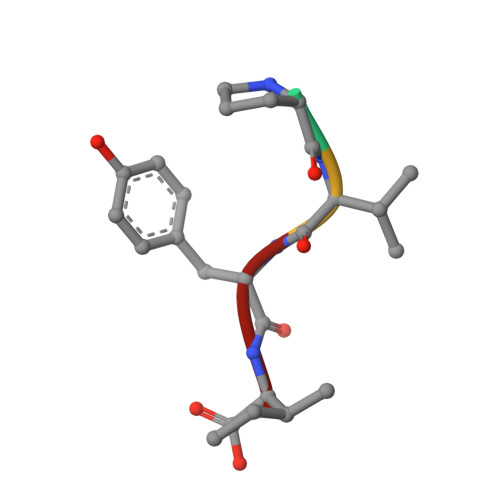
Autoinhibition of X11/Mint scaffold proteins revealed by the closed conformation of the PDZ tandem
Long, J.-F., Feng, W., Wang, R., Chan, L.-N., Ip, F.C., Xia, J., Ip, N.Y., Zhang, M.(2005) Nat Struct Mol Biol 12: 722-728
- PubMed: 16007100 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb958
- Primary Citation Related Structures:
1U37, 1U38, 1U39, 1U3B - PubMed Abstract:
Members of the X11/Mint family of multidomain adaptor proteins are composed of a divergent N terminus, a conserved PTB domain and a pair of C-terminal PDZ domains. Many proteins can interact with the PDZ tandem of X11 proteins, although the mechanism of such interactions is unclear. Here we show that the highly conserved C-terminal tail of X11alpha folds back and inserts into the target-binding groove of the first PDZ domain. The binding of this tail occludes the binding of other target peptides. This autoinhibited conformation of X11 requires that the two PDZ domains and the entire C-terminal tail be covalently connected to form an integral structural unit. The autoinhibited conformation of the X11 PDZ tandem provides a mechanistic explanation for the unique target-binding properties of the protein and hints at potential regulatory mechanisms for the X11-target interactions.
- Department of Biochemistry, Molecular Neuroscience Center, Hong Kong University of Science and Technology, Clear Water Bay, Kowloon, Hong Kong, China.
Organizational Affiliation: