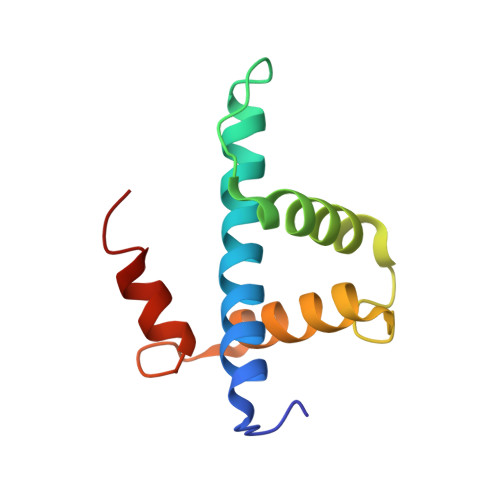
CBP/p300 TAZ1 domain forms a structured scaffold for ligand binding
De Guzman, R.N., Wojciak, J.M., Martinez-Yamout, M.A., Dyson, H.J., Wright, P.E.(2005) Biochemistry 44: 490-497
- PubMed: 15641773 Search on PubMed
- DOI: https://doi.org/10.1021/bi048161t
- Primary Citation Related Structures:
1U2N - PubMed Abstract:
The transcriptional coactivator protein CBP and its paralog p300 each contain two homologous zinc-containing TAZ domains, which constitute the interaction sites for a number of transcription factors. Previous reports of the three-dimensional structures of TAZ1 in complex with binding partners and of the isolated CBP TAZ2 domain show a distinctive topology composed of four amphipathic helices, organized by three zinc-binding clusters with HCCC-type coordination. The isolated CBP TAZ2 domain forms a stable three-dimensional structure in solution, but a recent report [Dial, R., Sun, Z., and Freedman, S. J. (2003) Biochemistry 42, 9937] suggested that the isolated p300 TAZ1 domain lacks a well-defined structure and behaves like a molten globule, even in the presence of Zn(2+), and that the formation of a stable three-dimensional structure requires binding of a protein partner. In marked contrast to this result, we find that both the CBP and p300 TAZ domains in the presence of stoichiometric concentrations of Zn(2+) adopt a well-defined structure in solution in the absence of binding partners. We have determined the three-dimensional structure of the isolated CBP TAZ1 domain by NMR methods and show that it has the same structure in the presence and absence of binding partners. This is an important finding: whether the free TAZ1 domain forms a folded structure or behaves as a molten globule will have a significant bearing on the mechanism of protein-protein recognition. Although TAZ1 and TAZ2 share many structural similarities, there is a major structural difference: the fourth helix is oriented in opposite directions in the TAZ1 and TAZ2 domains. The structure of the free TAZ1 domain suggests that this difference is an inherent feature that determines binding specificity and facilitates discrimination between different subsets of transcription factors by the two TAZ domains.
- Department of Molecular Biology and Skaggs Institute for Chemical Biology, The Scripps Research Institute, 10550 North Torrey Pines Road, La Jolla, California 92037, USA.
Organizational Affiliation: