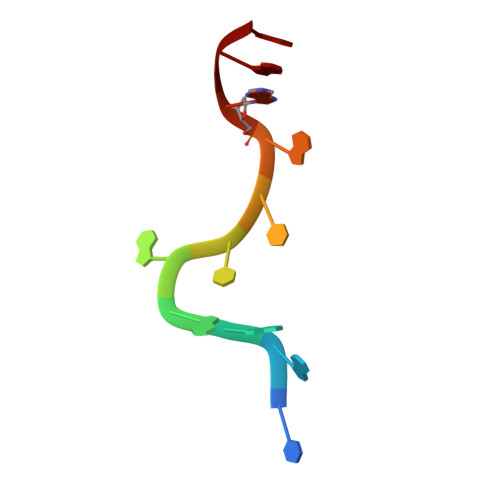
Human UP1 as a Model for Understanding Purine Recognition in the Family of Proteins Containing the RNA Recognition Motif (RRM).
Myers, J.C., Shamoo, Y.(2004) J Mol Biology 342: 743-756
- PubMed: 15342234 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.07.029
- Primary Citation Related Structures:
1U1K, 1U1L, 1U1M, 1U1N, 1U1O, 1U1P, 1U1Q, 1U1R - PubMed Abstract:
Heterogeneous ribonucleoprotein A1 (hnRNP A1) is a prototype for the family of eukaryotic RNA processing proteins containing the common RNA recognition motif (RRM). The region consisting of residues 1-195 of hnRNP A1 is referred to as UP1. This region has two RRMs and has a high affinity for both single-stranded RNA and the human telomeric repeat sequence d(TTAGGG)(n). We have used UP1's novel DNA binding to investigate how RRMs bind nucleic acid bases through their highly conserved RNP consensus sequences. Nine complexes of UP1 bound to modified telomeric repeats were investigated using equilibrium fluorescence binding and X-ray crystallography. In two of the complexes, alteration of a guanine to either 2-aminopurine or nebularine resulted in an increase in K(d) from 88nM to 209nM and 316nM, respectively. The loss of these orienting interactions between UP1 and the substituted base allows it to flip between syn and anti conformations. Substitution of the same base with 7-deaza-guanine preserves the O6/N1 contacts but still increases the K(d) to 296nM and suggests that it is not simply the loss of affinity that gives rise to the base mobility, but also the stereochemistry of the specific contact to O6. Although these studies provide details of UP1 interactions to nucleic acids, three general observations about RRMs are also evident: (1) as suggested by informatic studies, main-chain to base hydrogen bonding makes up an important aspect of ligand recognition (2) steric clashes generated by modification of a hydrogen bond donor-acceptor pair to a donor-donor pair are poorly tolerated and (3) a conserved lysine position proximal to RNP-2 (K(106)-IFVGGI) orients the purine to allow stereochemical discrimination between adenine and guanine based on the 6-position. This single interaction is well-conserved in known RRM structures and appears to be a broad indicator for purine preference in the larger family of RRM proteins.
- Department of Biochemistry and Cell Biology, Rice University, 6100 S. Main Street-MS140, Houston TX 77005, USA.
Organizational Affiliation: