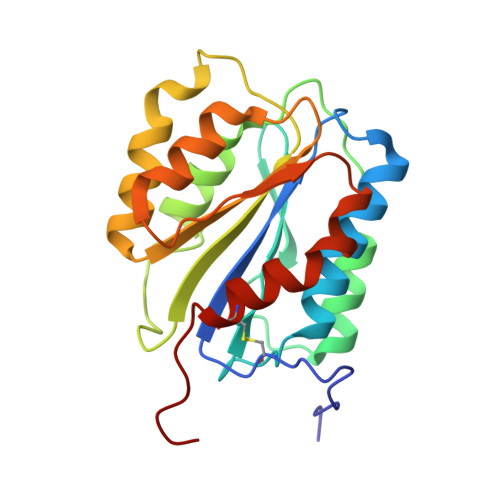
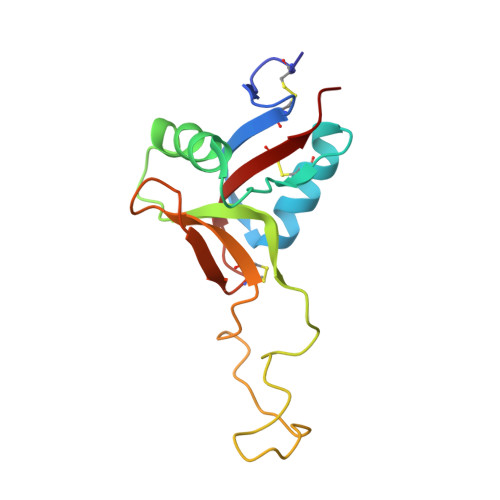
The snake venom protein botrocetin acts as a biological brace to promote dysfunctional platelet aggregation
Fukuda, K., Doggett, T., Laurenzi, I.J., Liddington, R.C., Diacovo, T.G.(2005) Nat Struct Mol Biol 12: 152-159
- PubMed: 15665869 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb892
- Primary Citation Related Structures:
1U0N, 1U0O - PubMed Abstract:
Botrocetin is a snake venom protein that enhances the affinity of the A1 domain of plasma von Willebrand factor (vWF) for the platelet receptor glycoprotein Ibalpha (GPIbalpha), an event that contributes to bleeding and host death. Here we describe a kinetic and crystallographic analysis of this interaction that reveals a novel mechanism of affinity enhancement. Using high-temporal-resolution microscopy, we show that botrocetin decreases the GPIbalpha off-rate two-fold in both human and mouse complexes without affecting the on-rate. The key to this behavior is that, upon binding of GPIbalpha to vWF-A1, botrocetin prebound to vWF-A1 makes no contacts initially with GPIbalpha, but subsequently slides around the A1 surface to form a new interface. This two-step mechanism and flexible coupling may prevent adverse alterations in on-rate of GPIbalpha for vWF-A1, and permit adaptation to structural differences in GPIbalpha and vWF in several prey species.
- Infectious and Inflammatory Disease Center, The Burnham Institute, 10901 North Torrey Pines Road, La Jolla, California 92037, USA.
Organizational Affiliation: