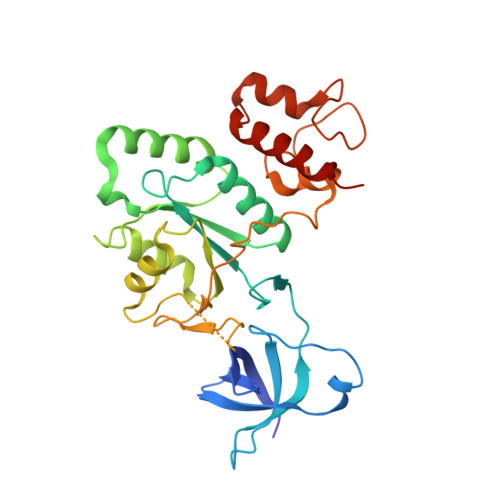
Crystal structure of YjeQ from Thermotoga maritima contains a circularly permuted GTPase domain
Shin, D.H., Lou, Y., Jancarik, J., Yokota, H., Kim, R., Kim, S.H.(2004) Proc Natl Acad Sci U S A 101: 13198-13203
- PubMed: 15331784 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.0405202101
- Primary Citation Related Structures:
1U0L - PubMed Abstract:
We have determined the crystal structure of the GDP complex of the YjeQ protein from Thermotoga maritima (TmYjeQ), a member of the YjeQ GTPase subfamaily. TmYjeQ, a homologue of Escherichia coli YjeQ, which is known to bind to the ribosome, is composed of three domains: an N-terminal oligonucleotide/oligosaccharide-binding fold domain, a central GTPase domain, and a C-terminal zinc-finger domain. The crystal structure of TmYjeQ reveals two interesting domains: a circularly permutated GTPase domain and an unusual zinc-finger domain. The binding mode of GDP in the GTPase domain of TmYjeQ is similar to those of GDP or GTP analogs in ras proteins, a prototype GTPase. The N-terminal oligonucleotide/oligosaccharide-binding fold domain, together with the GTPase domain, forms the extended RNA-binding site. The C-terminal domain has an unusual zinc-finger motif composed of Cys-250, Cys-255, Cys-263, and His-257, with a remote structural similarity to a portion of a DNA-repair protein, rad51 fragment. The overall structural features of TmYjeQ make it a good candidate for an RNA-binding protein, which is consistent with the biochemical data of the YjeQ subfamily in binding to the ribosome.
- Berkeley Structural Genomics Center, Lawrence Berkeley National Laboratory, Berkeley, CA 94720, USA.
Organizational Affiliation: