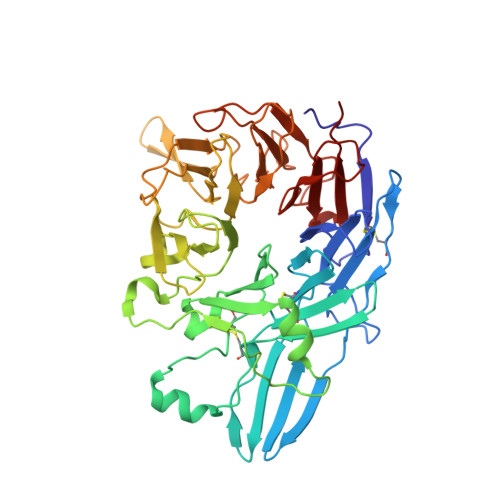
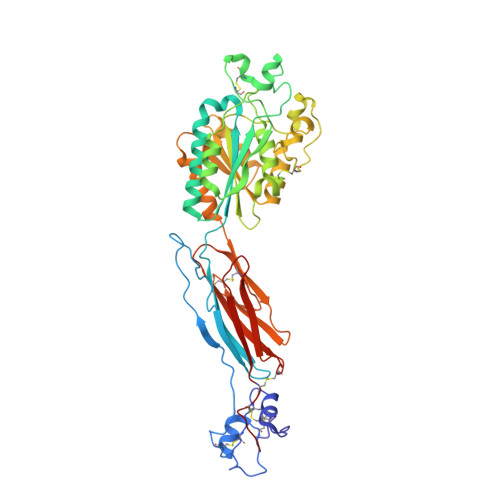
Structural basis for allostery in integrins and binding to fibrinogen-mimetic therapeutics
Xiao, T., Takagi, J., Coller, B.S., Wang, J.-H., Springer, T.A.(2004) Nature 432: 59-67
- PubMed: 15378069
- DOI: https://doi.org/10.1038/nature02976
- Primary Citation Related Structures:
1TYE - PubMed Abstract:
Integrins are important adhesion receptors in all Metazoa that transmit conformational change bidirectionally across the membrane. Integrin alpha and beta subunits form a head and two long legs in the ectodomain and span the membrane. Here, we define with crystal structures the atomic basis for allosteric regulation of the conformation and affinity for ligand of the integrin ectodomain, and how fibrinogen-mimetic therapeutics bind to platelet integrin alpha(IIb)beta3. Allostery in the beta3 I domain alters three metal binding sites, associated loops and alpha1- and alpha7-helices. Piston-like displacement of the alpha7-helix causes a 62 degrees reorientation between the beta3 I and hybrid domains. Transmission through the rigidly connected plexin/semaphorin/integrin (PSI) domain in the upper beta3 leg causes a 70 A separation between the knees of the alpha and beta legs. Allostery in the head thus disrupts interaction between the legs in a previously described low-affinity bent integrin conformation, and leg extension positions the high-affinity head far above the cell surface.
- The CBR Institute for Biomedical Research and Department of Pathology, Harvard Medical School, 200 Longwood Avenue, Boston, Massachusetts 02115, USA.
Organizational Affiliation: