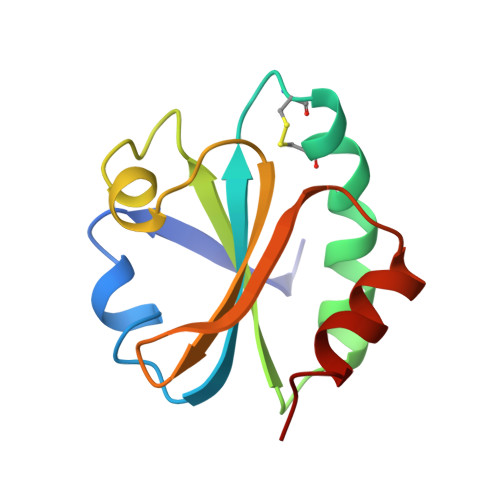
The CXXC motif: crystal structure of an active-site variant of Escherichia coli thioredoxin.
Schultz, L.W., Chivers, P.T., Raines, R.T.(1999) Acta Crystallogr D Biol Crystallogr 55: 1533-1538
- PubMed: 10489448 Search on PubMed
- DOI: https://doi.org/10.1107/s0907444999008756
- Primary Citation Related Structures:
1TXX - PubMed Abstract:
The 2.2 A crystalline structure of an oxidized active-site variant of Escherichia coli thioredoxin (Trx) has been solved. Trx is a 12 kDa enzyme which catalyzes the oxidation of dithiols and the reduction and isomerization of disulfides in other proteins. Its active site contains the common structural motif CXXC. Protein-disulfide isomerase (PDI), a 57 kDa homolog of Trx, contains four Trx-like domains. The three-dimensional structure of PDI is unknown. PDI-deficient Saccharomyces cerevisiae are inviable. An active-site variant of Trx which complements PDI-deficient yeast has the active-site sequence Cys32-Val33-Trp34-Cys35 (CVWC). The reduction potential of oxidized CVWC Trx (E degrees ' = -0.230 V) is altered significantly from that of the wild-type enzyme (E degrees ' = -0.270 V). However, the structure of the oxidized CVWC enzyme is almost identical to that of wild-type Trx. The addition of valine and tryptophan in the active site is likely to increase the reduction potential, largely by decreasing the pK(a) of the Cys32 thiol in the reduced enzyme. Unlike in wild-type Trx, significant protein-protein contacts occur in the crystal. Protein molecules related by a crystallographic twofold axis form a dimer in the crystal. The dimer forms as an extension of the twisted mixed beta-sheet which composes the backbone of each Trx structure.
- Department of Biochemistry, University of Wisconsin-Madison, Madison, WI 53706, USA.
Organizational Affiliation: