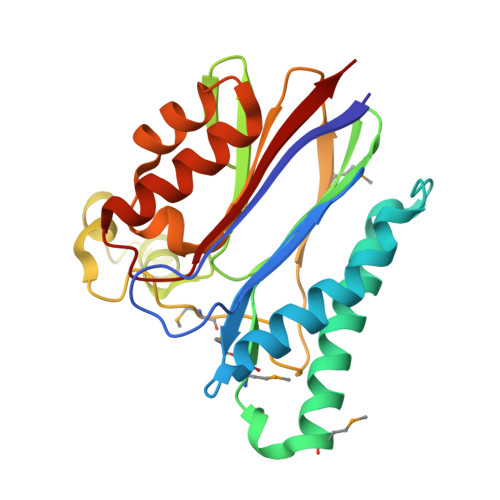
An Alternate Conformation and a Third Metal in PstP/Ppp, the M. tuberculosis PP2C-Family Ser/Thr Protein Phosphatase.
Pullen, K.E., Ng, H.L., Sung, P.Y., Good, M.C., Smith, S.M., Alber, T.(2004) Structure 12: 1947-1954
- PubMed: 15530359 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.09.008
- Primary Citation Related Structures:
1TXO - PubMed Abstract:
Serine/threonine protein phosphatases are central mediators of phosphorylation-dependent signals in eukaryotes and a variety of pathogenic bacteria. Here, we report the crystal structure of the intracellular catalytic domain of Mycobacterium tuberculosis PstPpp, a membrane-anchored phosphatase in the PP2C family. Despite sharing the fold and two-metal center of human PP2Calpha, the PstPpp catalytic domain binds a third Mn(2+) in a site created by a large shift in a previously unrecognized flap subdomain adjacent to the active site. Mutations in this site selectively increased the Michaelis constant for Mn(2+) in the reaction of a noncognate, small-molecule substrate, p-nitrophenyl phosphate. The PstP/Ppp structure reveals core functional motifs that advance the framework for understanding the mechanisms of substrate recognition, catalysis, and regulation of PP2C phosphatases.
- Department of Molecular and Cell Biology, 339 Hildebrand Hall #3206, University of California, Berkeley, Berkeley, CA 94720, USA.
Organizational Affiliation: