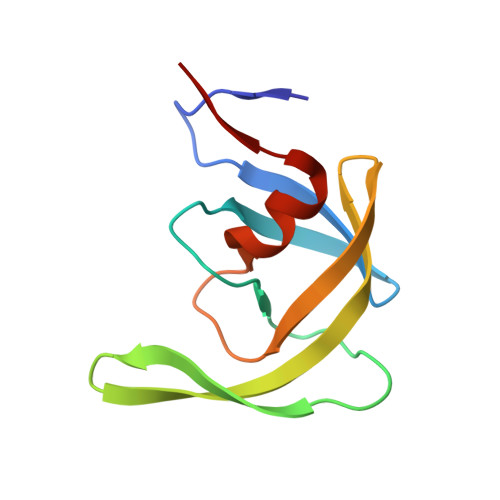
Wide Open 1.3A Structure of a Multi-drug Resistant HIV-1 Protease Represents a Novel Drug Target
Martin, P., Vickrey, J.F., Proteasa, G., Jimenez, Y.L., Wawrzak, Z., Winters, M.A., Merigan, T.C., Kovari, L.C.(2005) Structure 13: 1887-1895
- PubMed: 16338417 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2005.11.005
- Primary Citation Related Structures:
1TW7 - PubMed Abstract:
This report examines structural changes in a highly mutated, clinical multidrug-resistant HIV-1 protease, and the crystal structure has been solved to 1.3 A resolution in the absence of any inhibitor. This protease variant contains codon mutations at positions 10, 36, 46, 54, 62, 63, 71, 82, 84, and 90 that confer resistance to protease inhibitors. Major differences between the wild-type and the variant include a structural change initiated by the M36V mutation and amplified by additional mutations in the flaps of the protease, resulting in a "wide-open" structure that represents an opening that is 8 A wider than the "open" structure of the wild-type protease. A second structural change is triggered by the L90M mutation that results in reshaping the 23-32 segment. A third key structural change of the protease is due to the mutations from longer to shorter amino acid side chains at positions 82 and 84.
- Department of Biochemistry and Molecular Biology, Wayne State University School of Medicine, Detroit, Michigan 48201, USA.
Organizational Affiliation: