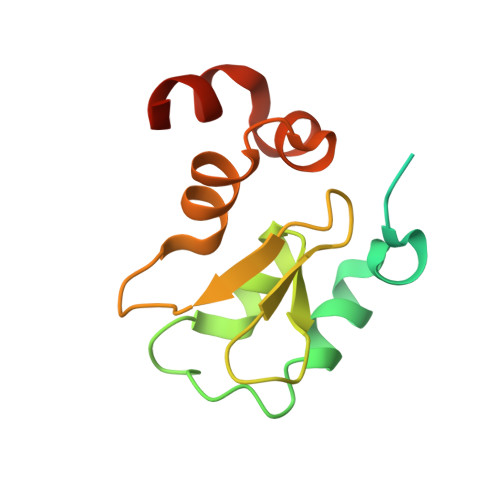
Engineering ML-IAP to produce an extraordinarily potent caspase 9 inhibitor: implications for Smac-dependent anti-apoptotic activity of ML-IAP
Vucic, D., Franklin, M.C., Wallweber, H.J.A., Das, K., Eckelman, B.P., Shin, H., Elliott, L.O., Kadkhodayan, S., Deshayes, K., Salvesen, G.S., Fairbrother, W.J.(2005) Biochem J 385: 11-20
- PubMed: 15485396 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BJ20041108
- Primary Citation Related Structures:
1TW6 - PubMed Abstract:
ML-IAP (melanoma inhibitor of apoptosis) is a potent anti-apoptotic protein that is strongly up-regulated in melanoma and confers protection against a variety of pro-apoptotic stimuli. The mechanism by which ML-IAP regulates apoptosis is unclear, although weak inhibition of caspases 3 and 9 has been reported. Here, the binding to and inhibition of caspase 9 by the single BIR (baculovirus IAP repeat) domain of ML-IAP has been investigated and found to be significantly less potent than the ubiquitously expressed XIAP (X-linked IAP). Engineering of the ML-IAP-BIR domain, based on comparisons with the third BIR domain of XIAP, resulted in a chimeric BIR domain that binds to and inhibits caspase 9 significantly better than either ML-IAP-BIR or XIAP-BIR3. Mutational analysis of the ML-IAP-BIR domain demonstrated that similar enhancements in caspase 9 affinity can be achieved with only three amino acid substitutions. However, none of these modifications affected binding of the ML-IAP-BIR domain to the IAP antagonist Smac (second mitochondrial activator of caspases). ML-IAP-BIR was found to bind mature Smac with low nanomolar affinity, similar to that of XIAP-BIR2-BIR3. Correspondingly, increased expression of ML-IAP results in formation of a ML-IAP-Smac complex and disruption of the endogenous interaction between XIAP and mature Smac. These results suggest that ML-IAP might regulate apoptosis by sequestering Smac and preventing it from antagonizing XIAP-mediated inhibition of caspases, rather than by direct inhibition of caspases.
- Department of Molecular Oncology, Genentech, Inc., 1 DNA Way, South San Francisco, CA 94080, USA.
Organizational Affiliation: