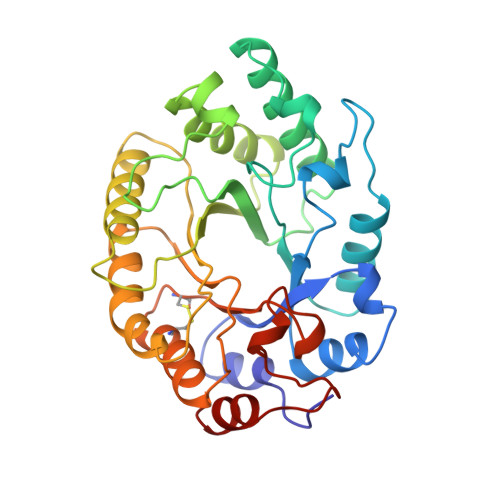
Crystal structure at 1.8 A resolution and proposed amino acid sequence of a thermostable xylanase from Thermoascus aurantiacus.
Natesh, R., Bhanumoorthy, P., Vithayathil, P.J., Sekar, K., Ramakumar, S., Viswamitra, M.A.(1999) J Mol Biology 288: 999-1012
- PubMed: 10329194 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1999.2727
- Primary Citation Related Structures:
1TUX - PubMed Abstract:
Thermoascus aurantiacus xylanase is a thermostable enzyme which hydrolyses xylan, a major hemicellulose component in the biosphere. Crystals belonging to P21 space group with a=41.7 A, b=68.1 A, c=51. 4 A and beta=113.6 degrees, Z=2 were grown that could diffract to better than 1.8 A resolution. The structure was solved by molecular replacement method using the Streptomyces lividans xylanase model. The amino acid sequence was determined from the electron density map aided by multiple alignment of related xylanase sequences. The sequence thus obtained provides a correction to the sequence reported earlier based on biochemical methods. The final refined protein model at 1.8 A resolution with 301 amino acid residues and 266 water molecules has an R-factor of 16.0 % and free R of 21.1 % with good stereochemistry. The single polypeptide chain assumes (alpha/beta)8 TIM-barrel fold and belongs to F/10 family of glycoside hydrolases. The active site consists of two glutamate residues located at the C terminus end of the beta-barrel, conforming to the double displacement mechanism for the enzyme action. A disulphide bond and more than ten salt bridges have been identified. In particular, the salt bridge Arg124-Glu232 which is almost buried, bridges the beta-strands beta4 and beta7 where the catalytic glutamate residues reside, and it may play a key role in the stability and activity at elevated temperature. To our knowledge, for the first time in the F/10 family xylanases, we observe a proline residue in the middle of the alpha-helix alpha6 which may be contributing to better packing. Earlier studies show that the enzyme retains its activity even at 70 degrees C. The refined protein model has allowed a detailed comparison with the other known structures in the F/10 family of enzymes. The possible causative factors for thermostability are discussed.
- Department of Physics, Indian Institute of Science, Bangalore, 560 012, India.
Organizational Affiliation: