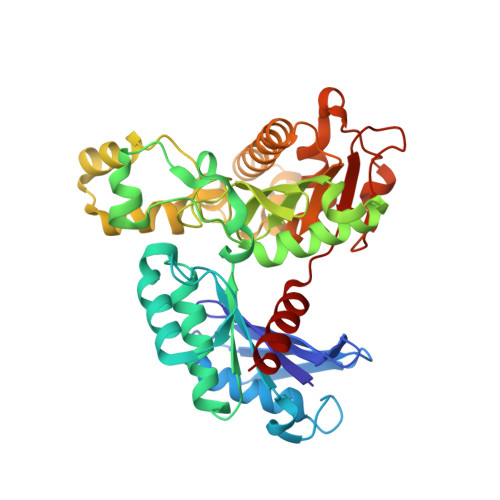
Structural and kinetic analyses of arginine residues in the active site of the acetate kinase from Methanosarcina thermophila.
Gorrell, A., Lawrence, S.H., Ferry, J.G.(2005) J Biological Chem 280: 10731-10742
- PubMed: 15647264 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M412118200
- Primary Citation Related Structures:
1TUU, 1TUY - PubMed Abstract:
Acetate kinase catalyzes transfer of the gamma-phosphate of ATP to acetate. The only crystal structure reported for acetate kinase is the homodimeric enzyme from Methanosarcina thermophila containing ADP and sulfate in the active site (Buss, K. A., Cooper, D. C., Ingram-Smith, C., Ferry, J. G., Sanders, D. A., and Hasson, M. S. (2001) J. Bacteriol. 193, 680-686). Here we report two new crystal structure of the M. thermophila enzyme in the presence of substrate and transition state analogs. The enzyme co-crystallized with the ATP analog adenosine 5'-[gamma-thio]triphosphate contained AMP adjacent to thiopyrophosphate in the active site cleft of monomer B. The enzyme co-crystallized with ADP, acetate, Al(3+), and F(-) contained a linear array of ADP-AlF(3)-acetate in the active site cleft of monomer B. Together, the structures clarify the substrate binding sites and support a direct in-line transfer mechanism in which AlF(3) mimics the meta-phosphate transition state. Monomers A of both structures contained ADP and sulfate, and the active site clefts were closed less than in monomers B, suggesting that domain movement contributes to catalysis. The finding that His(180) was in close proximity to AlF(3) is consistent with a role for stabilization of the meta-phosphate that is in agreement with a previous report indicating that this residue is essential for catalysis. Residue Arg(241) was also found adjacent to AlF(3), consistent with a role for stabilization of the transition state. Kinetic analyses of Arg(241) and Arg(91) replacement variants indicated that these residues are essential for catalysis and also indicated a role in binding acetate.
- Department of Biochemistry and Molecular Biology, Pennsylvania State University, University Park, Pennsylvania 16802, USA.
Organizational Affiliation: