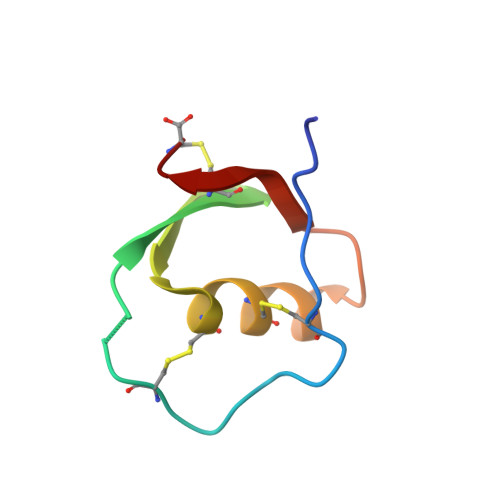
Solution structure of reactive-site hydrolyzed turkey ovomucoid third domain by nuclear magnetic resonance and distance geometry methods.
Walkenhorst, W.F., Krezel, A.M., Rhyu, G.I., Markley, J.L.(1994) J Mol Biology 242: 215-230
- PubMed: 8089843 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1994.1574
- Primary Citation Related Structures:
1TUS - PubMed Abstract:
The solution structure of reactive-site hydrolyzed turkey ovomucoid third domain (OMTKY3*) was determined by n.m.r. methods. A total of 655 distance constraints was applied in a distance geometry/simulated annealing approach to calculate a family of structures consistent with the n.m.r. data. The input data included 24 torsion angle constraints, 14 hydrogen bonds, 611 constraints derived from two-dimensional nuclear Overhauser enhancement spectroscopy data, and three disulfide bridges. Stereospecific assignments were included for the hydrogens of 26 beta-methylene groups and for seven isopropyl methyl groups (46% chiral assignments). OMTKY3* in solution retains the global fold and overall secondary structure of the intact inhibitor (OMTKY3) but exhibits local structural differences at and adjacent to the clip site. In particular, the hydrogen-bonding network observed at the reactive-site of the intact inhibitor is disrupted, and the position of Tyr20 is altered in the modified inhibitor. No evidence was found for ion pairing between the oppositely charged termini at the clip site. Surprisingly, in light of numerous changes indicating that OMTKY3* is less stable than OMTKY3, rotation of the Tyr31 ring was found to be slow in OMTKY3* at 30 degrees C. In OMTKY3, slow rotation of the Tyr31 ring was observed only at temperatures below 15 degrees C. The n.m.r. structures of OMTKY3* are compared here with the similarly calculated structures of OMTKY3. This represents the first comparison of an intact and modified (reactive-site clipped) proteinase inhibitor under identical conditions. On comparison with published X-ray structures of modified avian ovomucoid third domains from two other species, the present structure of OMTKY3* in solution was found to resemble that of the Japanese quail protein (OMJPQ3*) more closely than that of the more closely homologous silver pheasant protein (OMSVP3*).
- Department of Biochemistry, College of Agricultural and Life Sciences, University of Wisconsin-Madison 53706.
Organizational Affiliation: