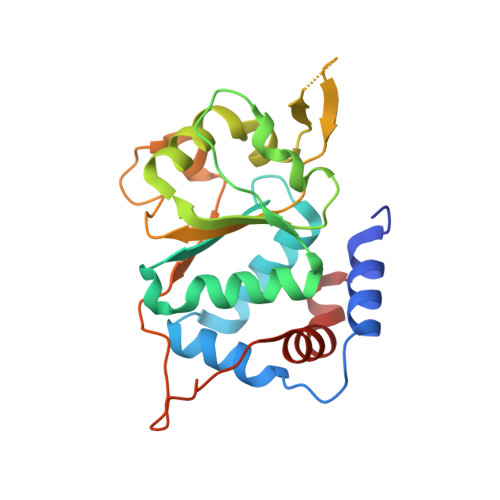
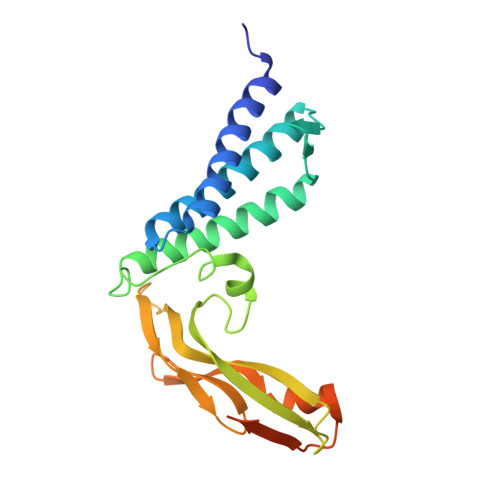
The X-ray structure of the papillomavirus helicase in complex with its molecular matchmaker E2
Abbate, E.A., Berger, J.M., Botchan, M.R.(2004) Genes Dev 18: 1981-1996
- PubMed: 15289463 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/gad.1220104
- Primary Citation Related Structures:
1TUE - PubMed Abstract:
DNA replication of the papillomaviruses is specified by cooperative binding of two proteins to the ori site: the enhancer E2 and the viral initiator E1, a distant member of the AAA+ family of proteins. Formation of this prereplication complex is an essential step toward the construction of a functional, multimeric E1 helicase and DNA melting. To understand how E2 interacts with E1 to regulate this process, we have solved the X-ray structure of a complex containing the HPV18 E2 activation domain bound to the helicase domain of E1. Modeling the monomers of E1 to a hexameric helicase shows that E2 blocks hexamerization of E1 by shielding a region of the E1 oligomerization surface and stabilizing a conformation of E1 that is incompatible with ATP binding. Further biochemical experiments and structural analysis show that ATP is an allosteric effector of the dissociation of E2 from E1. Our data provide the first molecular insights into how a protein can regulate the assembly of an oligomeric AAA+ complex and explain at a structural level why E2, after playing a matchmaker role by guiding E1 to the DNA, must dissociate for subsequent steps of initiation to occur. Building on previously proposed ideas, we discuss how our data advance current models for the conversion of E1 in the prereplication complex to a hexameric helicase assembly.
- Department of Molecular and Cell Biology, Division of Biochemistry and Molecular Biology, University of California, Berkeley, 94720-3204, USA.
Organizational Affiliation: