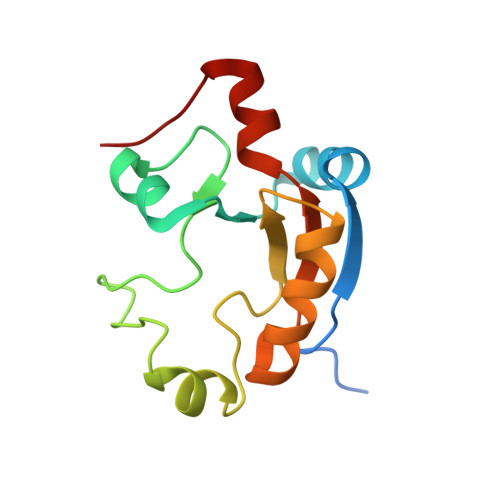
Solution structure of a single-domain thiosulfate sulfurtransferase from Arabidopsis thaliana.
Cornilescu, G., Vinarov, D.A., Tyler, E.M., Markley, J.L., Cornilescu, C.C.(2006) Protein Sci 15: 2836-2841
- PubMed: 17088324 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.062395206
- Primary Citation Related Structures:
1TQ1 - PubMed Abstract:
We describe the three-dimensional structure of the product of Arabidopsis thaliana gene At5g66040.1 as determined by NMR spectroscopy. This protein is categorized as single-domain sulfurtransferase and is annotated as a senescence-associated protein (sen1-like protein) and ketoconazole resistance protein (http://arabidopsis.org/info/genefamily/STR_genefamily.html). The sequence of At5g66040.1 is virtually identical to that of a protein from Arabidopsis found by others to confer ketoconazole resistance in yeast. Comparison of the three-dimensional structure with those in the Protein Data Bank revealed that At5g66040.1 contains an additional mobile beta-hairpin not found in other rhodaneses that may function in binding specific substrates. This represents the first structure of a single-domain plant sulfurtransferase. The enzymatically active cysteine-containing domain belongs to the CDC25 class of phosphatases, sulfide dehydrogenases, and stress proteins such as senescence specific protein 1 in plants, PspE and GlpE in bacteria, and cyanide and arsenate resistance proteins. Versions of this domain that lack the active site cysteine are found in other proteins, such as phosphatases, ubiquitin hydrolases, and sulfuryltransferases.
- National Magnetic Resonance Facility at Madison, Biochemistry Department, University of Wisconsin-Madison, Madison, Wisconsin 53706-1544, USA. gabrielc@nmrfam.wisc.edu
Organizational Affiliation: