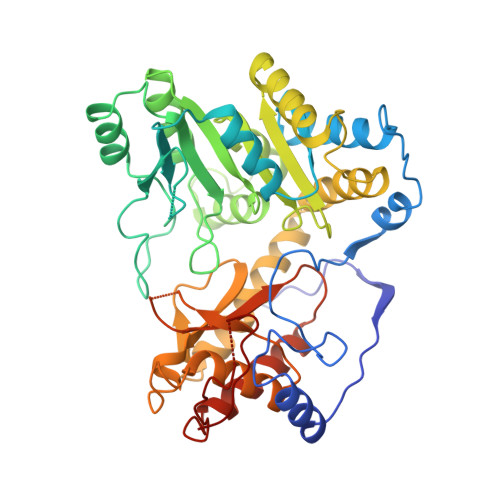
Three-dimensional structure of tyrosine phenol-lyase.
Antson, A.A., Demidkina, T.V., Gollnick, P., Dauter, Z., von Tersch, R.L., Long, J., Berezhnoy, S.N., Phillips, R.S., Harutyunyan, E.H., Wilson, K.S.(1993) Biochemistry 32: 4195-4206
- PubMed: 7916622 Search on PubMed
- DOI: https://doi.org/10.1021/bi00067a006
- Primary Citation Related Structures:
1TPL - PubMed Abstract:
Tyrosine phenol-lyase (EC 4.1.99.2) from Citrobacter freundii has been cloned and the primary sequence deduced from the DNA sequence. From the BrCN digest of the NaBH4-reduced holoenzyme, five peptides were purified and sequenced. The amino acid sequences of the peptides agreed with the corresponding parts of the tyrosine phenol-lyase sequence obtained from the gene structure. K257 is the pyridoxal 5'-phosphate binding residue. Assisted by the sequence data, the crystal structure of apotyrosine phenol-lyase, a pyridoxal 5'-phosphate-dependent enzyme, has been refined to an R-factor of 16.2% at 2.3-A resolution using synchrotron radiation diffraction data. The tetrameric molecule has 222 symmetry, with one of the axes coincident with the crystallographic 2-fold symmetry axis of the crystal which belongs to the space group P2(1)2(1)2 with a = 76.0 A, b = 138.3 A, and c = 93.5 A. Each subunit comprises 14 alpha-helices and 16 beta-strands, which fold into a small and a large domain. The coenzyme-binding lysine residue is located at the interface between the large and small domains of one subunit and the large domain of a crystallographically related subunit. The fold of the large, pyridoxal 5'-phosphate binding domain and the location of the active site are similar to that found in aminotransferases. Most of the residues which participate in binding of pyridoxal 5'-phosphate in aminotransferases are conserved in the structure of tyrosine phenol-lyase. Two dimers of tyrosine phenol-lyase, each of which has a domain architecture similar to that found in aspartate aminotransferases, are bound together through a hydrophobic cluster in the center of the molecule and intertwined N-terminal arms.
- Shubnikov Institute of Crystallography, Russian Academy of Sciences, Moscow.
Organizational Affiliation: