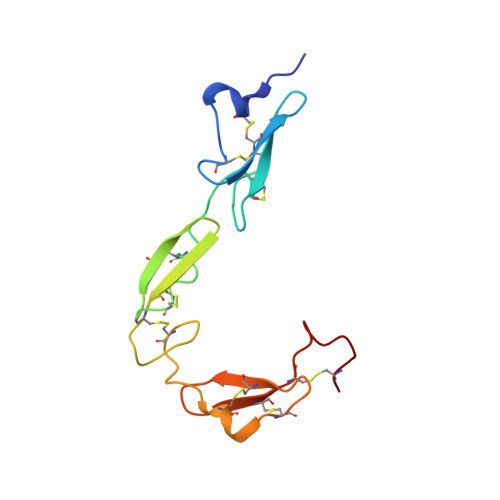
Structural and functional properties of the human notch-1 ligand binding region
Hambleton, S., Valeyev, N.V., Muranyi, A., Knott, V., Werner, J.M., McMichael, A.J., Handford, P.A., Downing, A.K.(2004) Structure 12: 2173-2183
- PubMed: 15576031 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.09.012
- Primary Citation Related Structures:
1TOZ - PubMed Abstract:
We present NMR structural and dynamics analysis of the putative ligand binding region of human Notch-1, comprising EGF-like domains 11-13. Functional integrity of an unglycosylated, recombinant fragment was confirmed by calcium-dependent binding of tetrameric complexes to ligand-expressing cells. EGF modules 11 and 12 adopt a well-defined, rod-like orientation rigidified by calcium. The interdomain tilt is similar to that found in previously studied calcium binding EGF pairs, but the angle of twist is significantly different. This leads to an extended double-stranded beta sheet structure, spanning the two EGF modules. Based on the conservation of residues involved in interdomain hydrophobic packing, we propose this arrangement to be prototypical of a distinct class of EGF linkages. On this premise, we have constructed a model of the 36 EGF modules of the Notch extracellular domain that enables predictions to be made about the general role of calcium binding to this region.
- Division of Molecular and Cellular Biochemistry, University of Oxford, South Parks Road, Oxford OX1 3QU, UK.
Organizational Affiliation: