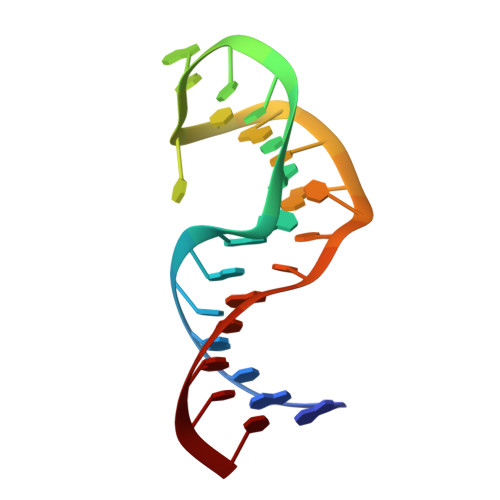
Saccharide-RNA recognition in an aminoglycoside antibiotic-RNA aptamer complex.
Jiang, L., Suri, A.K., Fiala, R., Patel, D.J.(1997) Chem Biol 4: 35
- PubMed: 9070426 Search on PubMed
- DOI: https://doi.org/10.1016/s1074-5521(97)90235-0
- Primary Citation Related Structures:
1TOB - PubMed Abstract:
Aminoglycoside antibiotics are known to target ribosomal, retroviral and catalytic RNAs with high affinity and specificity. Recently, in vitro selection experiments have identified RNA aptamers that bind to aminoglycoside antibiotics with nanomolar affinity and stringent specificity, allowing discrimination between closely related family members. There has, to date, been limited structural information on the molecular basis of such saccharide-RNA recognition. We describe a solution-structure determination of the tobramycin-RNA aptamer complex, obtained using NMR and molecular dynamics. The structure gives insight into the molecular features associated with saccharide-RNA recognition. Tobramycin adopts a defined alignment and binds to the RNA major groove centered about a stem-loop junction site. A portion of the bound tobramycin is encapsulated between the floor of the major groove and a looped-out cytosine residue that forms a flap over the binding site in the complex. The emergence of antibiotic-resistant pathogens and their impact on human health continues to be a major concern in the medical community. Rational modification of existing antibiotics aimed at improving their efficacy requires a molecular view of their receptor-binding sites. We have provided such a molecular view for a member of the aminoglycoside antibiotic family that targets RNA.
- Cellular Biochemistry and Biophysics Program, Memorial Sloan-Kettering Cancer Center, new York, NY 10021, USA.
Organizational Affiliation: