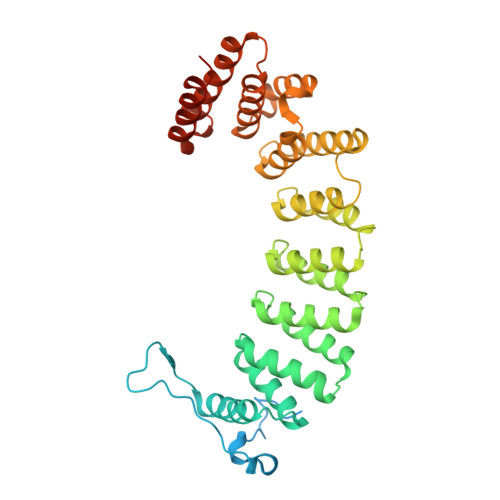
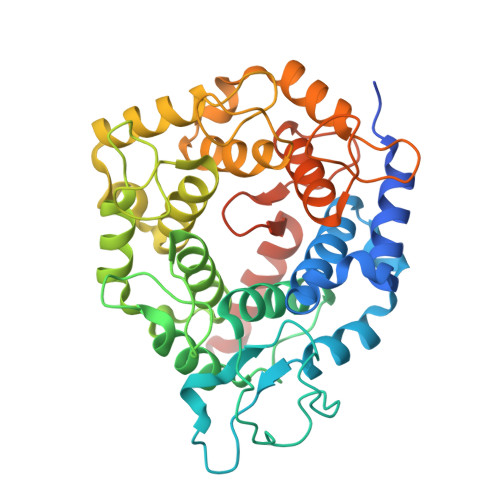
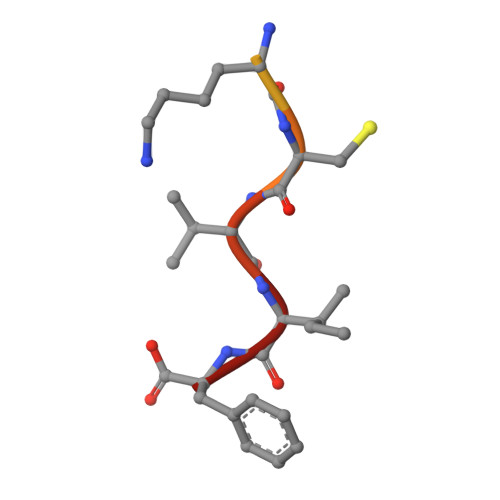
Crystallographic analysis of CaaX prenyltransferases complexed with substrates defines rules of protein substrate selectivity.
Reid, T.S., Terry, K.L., Casey, P.J., Beese, L.S.(2004) J Mol Biology 343: 417-433
- PubMed: 15451670 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.08.056
- Primary Citation Related Structures:
1TN6, 1TN7, 1TN8, 1TNB, 1TNO, 1TNU, 1TNY, 1TNZ - PubMed Abstract:
Post-translational modifications are essential for the proper function of many proteins in the cell. The attachment of an isoprenoid lipid (a process termed prenylation) by protein farnesyltransferase (FTase) or geranylgeranyltransferase type I (GGTase-I) is essential for the function of many signal transduction proteins involved in growth, differentiation, and oncogenesis. FTase and GGTase-I (also called the CaaX prenyltransferases) recognize protein substrates with a C-terminal tetrapeptide recognition motif called the Ca1a2X box. These enzymes possess distinct but overlapping protein substrate specificity that is determined primarily by the sequence identity of the Ca1a2X motif. To determine how the identity of the Ca1a2X motif residues and sequence upstream of this motif affect substrate binding, we have solved crystal structures of FTase and GGTase-I complexed with a total of eight cognate and cross-reactive substrate peptides, including those derived from the C termini of the oncoproteins K-Ras4B, H-Ras and TC21. These structures suggest that all peptide substrates adopt a common binding mode in the FTase and GGTase-I active site. Unexpectedly, while the X residue of the Ca1a2X motif binds in the same location for all GGTase-I substrates, the X residue of FTase substrates can bind in one of two different sites. Together, these structures outline a series of rules that govern substrate peptide selectivity; these rules were utilized to classify known protein substrates of CaaX prenyltransferases and to generate a list of hypothetical substrates within the human genome.
- Department of Biochemistry, Duke University Medical Center, Durham, NC 27710, USA.
Organizational Affiliation: