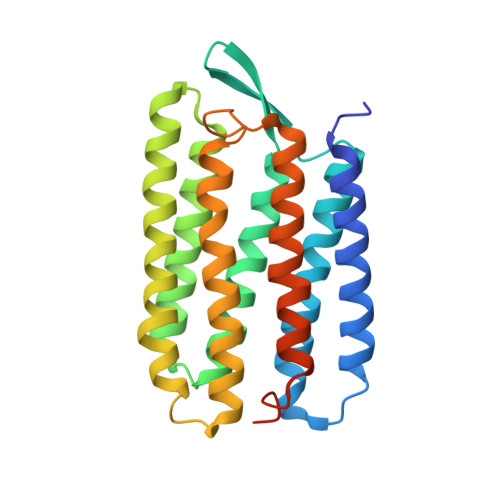
Proline substitutions are not easily accommodated in a membrane protein
Yohannan, S., Yang, D., Faham, S., Boulting, G., Whitelegge, J., Bowie, J.U.(2004) J Mol Biology 341: 1-6
- PubMed: 15312757 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2004.06.025
- Primary Citation Related Structures:
1TN0, 1TN5 - PubMed Abstract:
Proline residues are relatively common in transmembrane helices. This suggests that proline substitutions may be readily tolerated in membrane proteins, even though they invariably produce deviations from canonical helical structure. We have experimentally tested this possibility by making proline substitutions at 15 positions throughout the N-terminal half of bacteriorhodopsin helix B. We find that six of the substitutions yielded no active protein and all the others were destabilizing. Three mutations were only slightly destabilizing, however, reducing stability by about 0.5 kcal/mol, and these all occurred close to the N terminus. This result is consistent with the observation that proline is more common near the ends of TM helices. To learn how proline side-chains could be structurally accommodated at different locations in the helix, we solved the structures of a moderately destabilized mutant positioned near the N terminus of the helix, K41P, and a severely destabilized mutant positioned near the middle of the helix, A51P. The K41P mutation produced only local structural alterations, while the A51P mutation resulted in small, but widely distributed structural changes in helix B. Our results indicate that proline is not easily accommodated in transmembrane helices and that the tolerance to proline substitution is dependent, in a complex way, on the position in the structure.
- Department of Chemistry and Biochemistry, DOE Center for Genomics and Proteomics, Molecular Biology Institute, 655 Boyer Hall, 611 Charles E. Young Dr. E, University of California, Los Angeles, Los Angeles, CA 90095-1570, USA.
Organizational Affiliation: