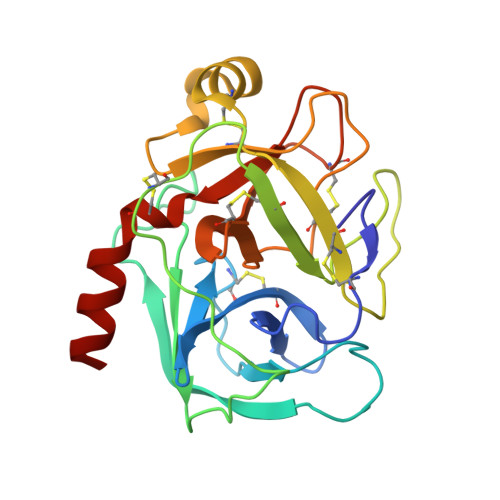
Crystal structure of bovine beta-trypsin at 1.5 A resolution in a crystal form with low molecular packing density. Active site geometry, ion pairs and solvent structure.
Bartunik, H.D., Summers, L.J., Bartsch, H.H.(1989) J Mol Biology 210: 813-828
- PubMed: 2614845 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(89)90110-1
- Primary Citation Related Structures:
1TLD - PubMed Abstract:
The crystal structure of bovine pancreatic beta-trypsin (BPT) has been determined from a novel orthorhombic crystal form which contains substantially more solvent (filling 57% of the volume of the unit cell) than previously determined orthorhombic (44%) and trigonal (37%) BPT structures. The native and benzamidine-inhibited crystal structures of BPT in ammonium sulphate at pH 5.3 have been determined for the new form by molecular replacement techniques. The structures have been refined at 1.5 A resolution with final R-values of 16.7% and 16.9%, respectively. Comparison with the previously refined old orthorhombic forms shows that the overall conformation of the protein backbone is highly conserved. A great number of previously undefined side-chains have been located in density. At the C terminus an extra ion pair involving lysines 87 and 107 has been revealed. A far more detailed picture of the ordered solvent structure has been derived. Thirty water clusters have been identified. A large water network extends from the calcium binding site to the activation area and the autolysis loop. There is evidence for a water channel reaching from the depth of the specificity pocket to the nearby protein surface which might be involved in the displacement of water molecules upon substrate binding. A sulphate anion which forms hydrogen bonds to the active site residues His57, Ser195 and Gly193 was for the first time positioned in clearly defined electron density. Interaction with the sulphate ion may explain the increase in the pKa value of His57 at high sulphate concentrations which was observed by nuclear magnetic resonance studies of a bacterial serine protease both in crystalline form and in solution. Thus, a His-Ser hydrogen bond will not exist in solvents containing sulphate at low pH (up to at least 6.8) where the imidazole of His57 is protonated. The new crystal form is of considerable interest for substrate binding studies. Wide solvent channels should allow diffusion of large substrates (comparable in size to, e.g. pancreatic trypsin inhibitor) into the enzyme crystal. The active site is accessible; intermolecular contact areas are further remote from the active site than in the old orthorhombic form.
- Max-Planck-Society, Research Unit for Structural Molecular Biology, Hamburg, West Germany.
Organizational Affiliation: