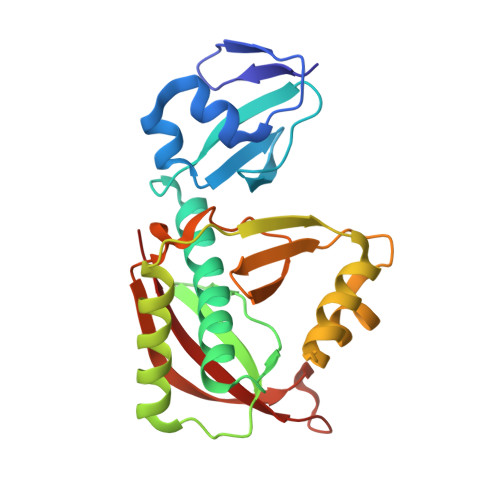
Achieving Error-Free Translation; The Mechanism of Proofreading of Threonyl-tRNA Synthetase at Atomic Resolution.
Dock-Bregeon, A.C., Rees, B., Torres-Larios, A., Bey, G., Caillet, J., Moras, D.(2004) Mol Cell 16: 375-386
- PubMed: 15525511
- DOI: https://doi.org/10.1016/j.molcel.2004.10.002
- Primary Citation of Related Structures:
1TJE, 1TKE, 1TKG, 1TKY - PubMed Abstract:
The fidelity of aminoacylation of tRNA(Thr) by the threonyl-tRNA synthetase (ThrRS) requires the discrimination of the cognate substrate threonine from the noncognate serine. Misacylation by serine is corrected in a proofreading or editing step. An editing site has been located 39 A away from the aminoacylation site. We report the crystal structures of this editing domain in its apo form and in complex with the serine product, and with two nonhydrolyzable analogs of potential substrates: the terminal tRNA adenosine charged with serine, and seryl adenylate. The structures show how serine is recognized, and threonine rejected, and provide the structural basis for the editing mechanism, a water-mediated hydrolysis of the mischarged tRNA. When the adenylate analog binds in the editing site, a phosphate oxygen takes the place of one of the catalytic water molecules, thereby blocking the reaction. This rules out a correction mechanism that would occur before the binding of the amino acid on the tRNA.
- IGBMC (CNRS/INSERM/Université Louis Pasteur), Laboratoire de Biologie et Génomique Structurales, 1, rue Laurent Fries, BP 10142, 67400 Illkirch, France.
Organizational Affiliation: