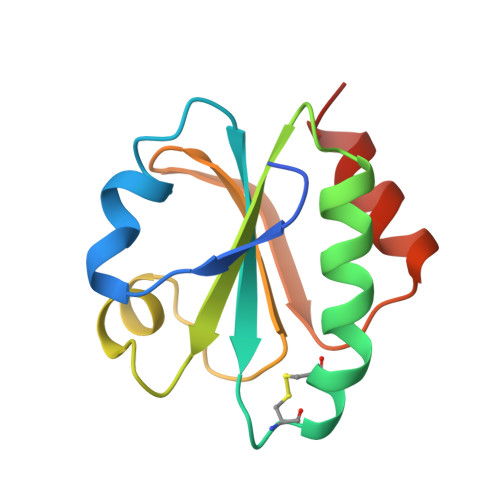
Crystal structure of thioredoxin-2 from Anabaena.
Saarinen, M., Gleason, F.K., Eklund, H.(1995) Structure 3: 1097-1108
- PubMed: 8590004 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(01)00245-3
- Primary Citation Related Structures:
1THX - PubMed Abstract:
Thioredoxins are ubiquitous proteins that serve as reducing agents and general protein disulfide reductases. The structures of thioredoxins from a number of species, including man and Escherichia coli, are known. Cyanobacteria, such as Anabaena, contain two thioredoxins that exhibit very different activities with target enzymes and share little sequence similarity. Thioredoxin-2 (Trx-2) from Anabaena resembles chloroplast type-f thioredoxin in its activities and the two proteins may be evolutionarily related. We have undertaken structural studies of Trx-2 in order to gain insights into the structure/function relationships of thioredoxins. Anabaena Trx-2, like E. coli thioredoxin, consists of a five-stranded beta sheet core surrounded by four alpha helices. The active site includes a conserved disulfide ring (in the sequence 31WCGPC35). An aspartate (E. coli) to tyrosine (Trx-2) substitution alters the position of this disulfide ring relative to the central pleated sheet. However, loss of this conserved aspartate does not affect the disulfide geometry. In the Trx-2 crystals, the N-terminal residues make extensive contacts with a symmetry-related molecule with hydrogen bonds to residues 74-76 mimicking thioredoxin-protein interactions. The overall three-dimensional structure of Trx-2 is similar to E. coli thioredoxin and other related disulfide oxido-reductases. Single amino acid substitutions around the protein interaction area probably account for the unusual enzymatic activities of Trx-2 and its ability to discriminate between substrate and non-substrate peptides.
- Department of Molecular Biology, Swedish University of Agricultural Sciences, Uppsala, Sweden.
Organizational Affiliation: