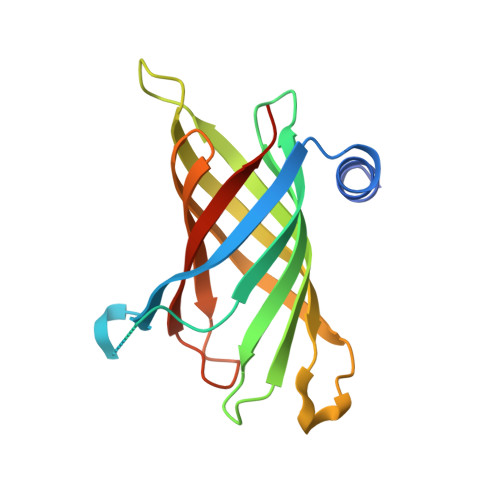
A hydrocarbon ruler measures palmitate in the enzymatic acylation of endotoxin.
Ahn, V.E., Lo, E.I., Engel, C.K., Chen, L., Hwang, P.M., Kay, L.E., Bishop, R.E., Prive, G.G.(2004) EMBO J 23: 2931-2941
- PubMed: 15272304 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/sj.emboj.7600320
- Primary Citation Related Structures:
1THQ - PubMed Abstract:
The ability of enzymes to distinguish between fatty acyl groups can involve molecular measuring devices termed hydrocarbon rulers, but the molecular basis for acyl-chain recognition in any membrane-bound enzyme remains to be defined. PagP is an outer membrane acyltransferase that helps pathogenic bacteria to evade the host immune response by transferring a palmitate chain from a phospholipid to lipid A (endotoxin). PagP can distinguish lipid acyl chains that differ by a single methylene unit, indicating that the enzyme possesses a remarkably precise hydrocarbon ruler. We present the 1.9 A crystal structure of PagP, an eight-stranded beta-barrel with an unexpected interior hydrophobic pocket that is occupied by a single detergent molecule. The buried detergent is oriented normal to the presumed plane of the membrane, whereas the PagP beta-barrel axis is tilted by approximately 25 degrees. Acyl group specificity is modulated by mutation of Gly88 lining the bottom of the hydrophobic pocket, thus confirming the hydrocarbon ruler mechanism for palmitate recognition. A striking structural similarity between PagP and the lipocalins suggests an evolutionary link between these proteins.
- Department of Medical Biophysics, University of Toronto, Ontario, Canada.
Organizational Affiliation: