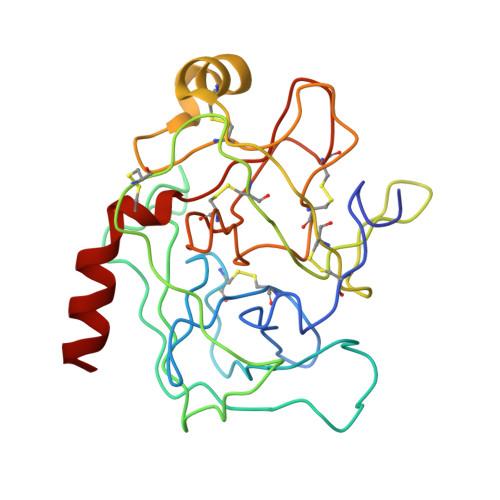
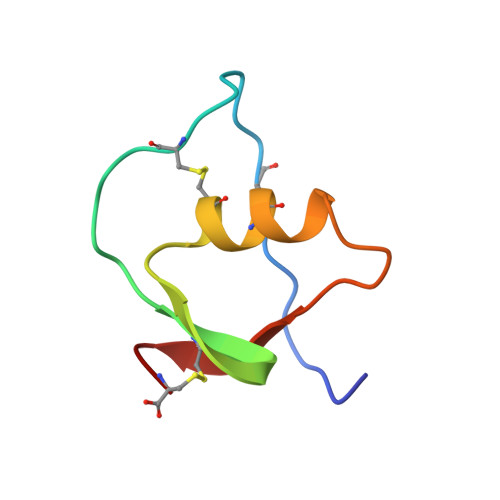
Three-dimensional structure of the complex between pancreatic secretory trypsin inhibitor (Kazal type) and trypsinogen at 1.8 A resolution. Structure solution, crystallographic refinement and preliminary structural interpretation.
Bolognesi, M., Gatti, G., Menagatti, E., Guarneri, M., Marquart, M., Papamokos, E., Huber, R.(1982) J Mol Biology 162: 839-868
- PubMed: 7169635 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(82)90550-2
- Primary Citation Related Structures:
1TGS