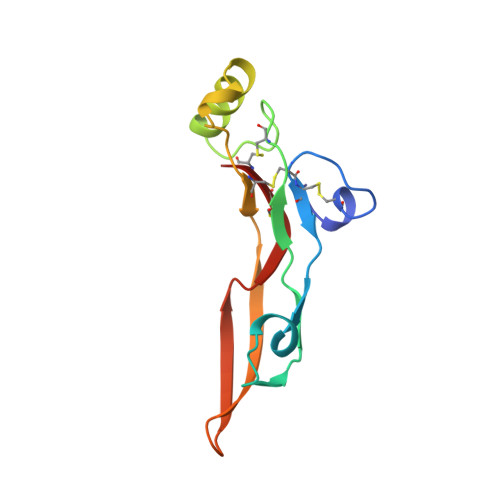
An unusual feature revealed by the crystal structure at 2.2 A resolution of human transforming growth factor-beta 2.
Schlunegger, M.P., Grutter, M.G.(1992) Nature 358: 430-434
- PubMed: 1641027 Search on PubMed
- DOI: https://doi.org/10.1038/358430a0
- Primary Citation Related Structures:
1TFG - PubMed Abstract:
Transforming growth factor type beta 2 (TGF-beta 2) is a member of an expanding family of growth factors that regulate proliferation and differentiation of many different cell types. TGF-beta 2 binds to various receptors, one of which was shown to be a serine/threonine kinase. TGF-beta 2 is involved in wound healing, bone formation and modulation of immune functions. We report here the crystal structure of TGF-beta 2 at 2.2 A resolution, which reveals a novel monomer fold and dimer association. The monomer consists of two antiparallel pairs of beta-strands forming a flat curved surface and a separate, long alpha-helix. The disulphide-rich core has one disulphide bone pointing through a ring formed by the sequence motifs Cys-Ala-Gly-Ala-Cys and Cys-Lys-Cys, which are themselves connected through the cysteines. Two monomers are connected through a single disulphide bridge and associate such that the helix of one subunit interacts with the concave beta-sheet surface of the other. Four exposed loop regions might determine receptor specificity. The structure provides a suitable model for the TGF-beta s and other members of the super-family and is the basis for the analysis of the TGF-beta 2 interactions with the receptor.
- Department of Biotechnology, Pharmaceuticals Division, Ciba-Geigy, Basel, Switzerland.
Organizational Affiliation: