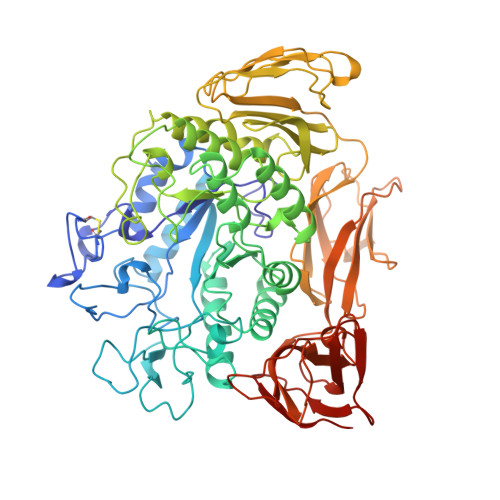
The raw starch binding domain of cyclodextrin glycosyltransferase from Bacillus circulans strain 251.
Penninga, D., van der Veen, B.A., Knegtel, R.M., van Hijum, S.A., Rozeboom, H.J., Kalk, K.H., Dijkstra, B.W., Dijkhuizen, L.(1996) J Biological Chem 271: 32777-32784
- PubMed: 8955113 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.271.51.32777
- Primary Citation Related Structures:
1TCM - PubMed Abstract:
The E-domain of cyclodextrin glycosyltransferase (CGTase) (EC 2.4.1.19) from Bacillus circulans strain 251 is a putative raw starch binding domain. Analysis of the maltose-dependent CGTase crystal structure revealed that each enzyme molecule contained three maltose molecules, situated at contact points between protein molecules. Two of these maltoses were bound to specific sites in the E-domain, the third maltose was bound at the C-domain. To delineate the roles in raw starch binding and cyclization reaction kinetics of the two maltose binding sites in the E-domain, we replaced Trp-616 and Trp-662 of maltose binding site 1 and Tyr-633 of maltose binding site 2 by alanines using site-directed mutagenesis. Purified mutant CGTases were characterized with respect to raw starch binding and cyclization reaction kinetics on both soluble and raw starch. The results show that maltose binding site 1 is most important for raw starch binding, whereas maltose binding site 2 is involved in guiding linear starch chains into the active site. beta-Cyclodextrin causes product inhibition by interfering with catalysis in the active site and the function of maltose binding site 2 in the E-domain. CGTase mutants in the E-domain maltose binding site 1 could no longer be crystallized as maltose-dependent monomers. Instead, the W616A mutant CGTase protein was successfully crystallized as a carbohydrate-independent dimer; its structure has been refined to 2.2 A resolution. The three-dimensional structure shows that, within the error limits, neither the absence of carbohydrates nor the W616A mutation caused significant further conformational changes. The modified starch binding and cyclization kinetic properties observed with the mutant CGTase proteins thus can be directly related to the amino acid replacements.
- Department of Microbiology, Groningen Biomolecular Sciences and Biotechnology Institute (GBB), University of Groningen, Kerklaan 30, 9751 NN Haren, The Netherlands. L.Dijkhuizen@biol.rug.nl
Organizational Affiliation: