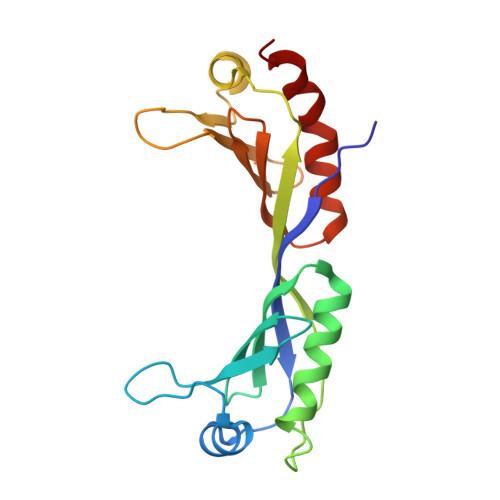
Crystal structure of yeast TATA-binding protein and model for interaction with DNA.
Chasman, D.I., Flaherty, K.M., Sharp, P.A., Kornberg, R.D.(1993) Proc Natl Acad Sci U S A 90: 8174-8178
- PubMed: 8367480 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.90.17.8174
- Primary Citation Related Structures:
1TBP - PubMed Abstract:
The C-terminal 179-aa region of yeast (Saccharomyces cerevisiae) TATA-binding protein (TBP), phylogenetically conserved and sufficient for many functions, formed crystals diffracting to 1.7-A resolution. The structure of the protein, determined by molecular replacement with coordinates from Arabidopsis TBP and refined to 2.6 A, differed from that in Arabidopsis slightly by an angle of about 12 degrees between two structurally nearly identical subdomains, indicative of a degree of conformational flexibility. A model for TBP-DNA interaction is proposed with the following important features: the long dimension of the protein follows the trajectory of the minor groove; two rows of basic residues conserved between the subdomains lie along the edges of the protein in proximity to the DNA phosphates; a band of hydrophobic residues runs down the middle of the groove; and amino acid residues whose mutation alters specificity for the second base of the TATA sequence are juxtaposed to that base.
- Center for Cancer Research, Massachusetts Institute of Technology, Cambridge 02139.
Organizational Affiliation: