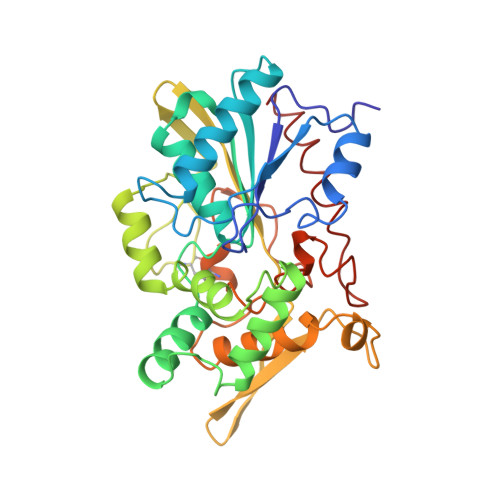
The crystal structure of triacylglycerol lipase from Pseudomonas glumae reveals a partially redundant catalytic aspartate.
Noble, M.E., Cleasby, A., Johnson, L.N., Egmond, M.R., Frenken, L.G.(1993) FEBS Lett 331: 123-128
- PubMed: 8405390 Search on PubMed
- DOI: https://doi.org/10.1016/0014-5793(93)80310-q
- Primary Citation Related Structures:
1TAH - PubMed Abstract:
The family of lipases (triacylglycerol-acyl-hydrolases EC 3.1.1.3) constitutes an interesting class of enzymes because of their ability to interact with lipid-water interfaces, their wide range of substrate specificities, and their potential industrial applications. Here we report the first crystal structure of a bacterial lipase, from Pseudomonas glumae. The structure is formed from three domains, the largest of which contains a subset of the alpha/beta hydrolase fold and a calcium site. Asp263, the acidic residue in the catalytic triad, has previously been mutated into an alanine with only a modest reduction in activity.
- Laboratory of Molecular Biophysics, Department of Biochemistry, Oxford, UK.
Organizational Affiliation: