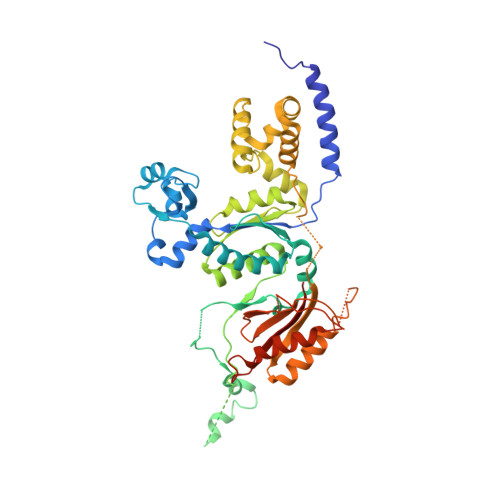
Crystal structure of the catalytic core of human DNA polymerase kappa.
Uljon, S.N., Johnson, R.E., Edwards, T.A., Prakash, S., Prakash, L., Aggarwal, A.K.(2004) Structure 12: 1395-1404
- PubMed: 15296733 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2004.05.011
- Primary Citation Related Structures:
1T94 - PubMed Abstract:
We present the crystal structure of the catalytic core of human DNA polymerase kappa (hPolkappa), the first structure of a human Y-family polymerase. hPolkappa is implicated in the proficient extension of mispaired primer termini on undamaged DNAs, and in the extension step of lesion bypass. The structure reveals a stubby "fingers" subdomain, which despite its small size appears to be tightly restrained with respect to a putative templating base. The structure also reveals a novel "thumb" subdomain that provides a basis for the importance of the N-terminal extension unique to hPolkappa. And, most surprisingly, the structure reveals the polymerase-associated domain (PAD) juxtaposed on the dorsal side of the "palm" subdomain, as opposed to the fingers subdomain. Together, these properties suggest that the hPolkappa active site is constrained at the site of the templating base and incoming nucleotide, but the polymerase is less constrained following translocation of the lesion.
- Structural Biology Program, Department of Physiology and Biophysics, Mount Sinai School of Medicine, Box 1677, 1425 Madison Avenue, New York, New York 10029, USA.
Organizational Affiliation: