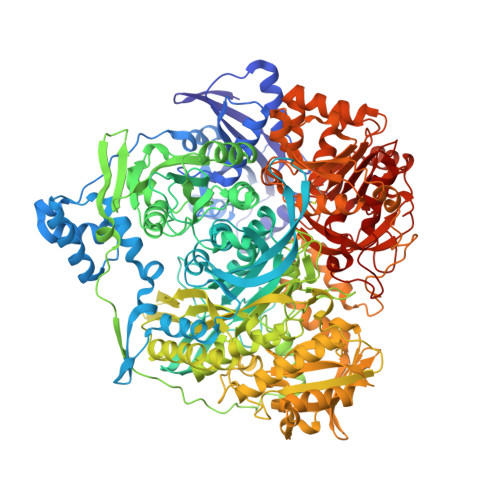
Domain Organization of Salmonella typhimurium Formylglycinamide Ribonucleotide Amidotransferase Revealed by X-ray crystallography
Anand, R., Hoskin, A.A., Stubbe, J., Ealick, S.E.(2004) Biochemistry 43: 10328-10342
- PubMed: 15301531 Search on PubMed
- DOI: https://doi.org/10.1021/bi0491301
- Primary Citation Related Structures:
1T3T - PubMed Abstract:
Formylglycinamide ribonucleotide amidotransferase (FGAR-AT) catalyzes the ATP-dependent conversion of formylglycinamide ribonucleotide (FGAR) and glutamine to formylglycinamidine ribonucleotide (FGAM), ADP, P(i), and glutamate in the fourth step of the purine biosynthetic pathway. In eukaryotes and Gram-negative bacteria, FGAR-AT is encoded by the purL gene as a multidomain protein with a molecular mass of about 140 kDa. In Gram-positive bacteria and archaebacteria FGAR-AT is a complex of three proteins: PurS, PurL, and PurQ. We have determined the structure of FGAR-AT (PurL) from Salmonella typhimurium at 1.9 A resolution using X-ray crystallography. PurL is the last remaining enzyme in the purine biosynthetic pathway to have its structure determined. The structure reveals four domains: an N-terminal domain structurally homologous to a PurS dimer, a linker region, an FGAM synthetase domain homologous to an aminoimidazole ribonucleotide synthetase (PurM) dimer, and a triad glutaminase domain. The domains are intricately linked by interdomain interactions and peptide connectors. The fold common to PurM and the central region of PurL represents a superfamily for which HypE, SelD, and ThiL are predicted to be members. A structural ADP molecule was found bound to a site related to the putative active site by pseudo-2-fold symmetry and two sulfate ions were found at the putative active site. These observations and the structural similarities between PurM and StPurL were used to model the substrates FGAR and ATP in the StPurL active site. A glutamylthioester intermediate was found in the glutaminase domain at Cys1135. The N-terminal (PurS-like) domain is hypothesized to form the putative channel through which ammonia passes from the glutaminase domain to the FGAM synthetase domain.
- Department of Chemistry and Chemical Biology, Cornell University, Ithaca, New York 14853, USA.
Organizational Affiliation: