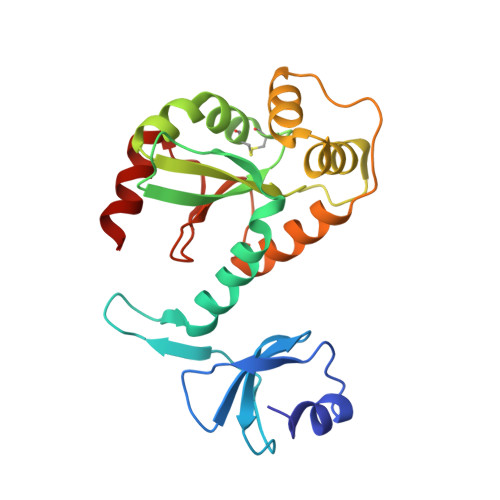
Structure of DsbC from Haemophilus influenzae.
Zhang, M., Monzingo, A.F., Segatori, L., Georgiou, G., Robertus, J.D.(2004) Acta Crystallogr D Biol Crystallogr 60: 1512-1518
- PubMed: 15333920 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444904014593
- Primary Citation Related Structures:
1T3B - PubMed Abstract:
Bacterial DsbC proteins are involved in rearranging or reducing mismatched disulfide bonds folding within the periplasm. The X-ray structure of the enzyme from Haemophilus influenzae has been solved and compared with the known structure of the Escherichia coli protein. The proteins act as V-shaped dimers with a large cleft to accommodate substrate proteins. The dimers are anchored by a small N-terminal domain, but have a flexible linker region which allows the larger C-terminal domain, with its reactive sulfhydryls, to clamp down on substrates. The overall folds are very similar, but the comparison shows a wider range of hinge motions than previously thought. The crystal packing of the H. influenzae protein allows the movement of the N-terminal domain with respect to the C-terminal domain through motions in the flexible hinge, generating high thermal parameters and unusually high anisotropy in the crystallographic data.
- Institute for Cellular and Molecular Biology, Department of Chemistry and Biochemistry, 1 University Station, University of Texas at Austin, Austin, TX 78712, USA.
Organizational Affiliation: