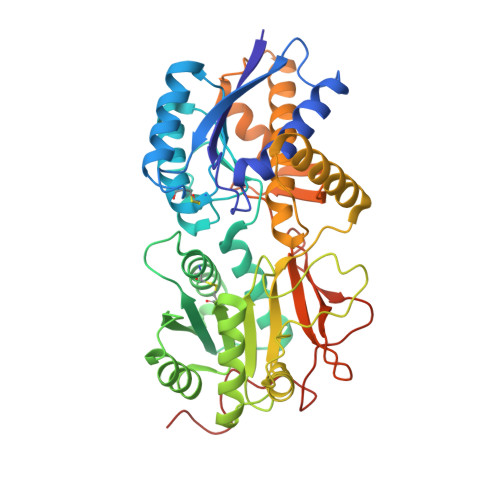
Crystal structure of hormone-bound atrial natriuretic peptide receptor extracellular domain: rotation mechanism for transmembrane signal transduction
Ogawa, H., Qiu, Y., Ogata, C.M., Misono, K.S.(2004) J Biological Chem 279: 28625-28631
- PubMed: 15117952 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M313222200
- Primary Citation Related Structures:
1T34 - PubMed Abstract:
A cardiac hormone, atrial natriuretic peptide (ANP), plays a major role in blood pressure and volume regulation. ANP activities are mediated by a single span transmembrane receptor carrying intrinsic guanylate cyclase activity. ANP binding to its extracellular domain stimulates guanylate cyclase activity by an as yet unknown mechanism. Here we report the crystal structure of dimerized extracellular hormone-binding domain in complex with ANP. The structural comparison with the unliganded receptor reveals that hormone binding causes the two receptor monomers to undergo an intermolecular twist with little intramolecular conformational change. This motion produces a Ferris wheel-like translocation of two juxtamembrane domains in the dimer with essentially no change in the interdomain distance. This movement alters the relative orientation of the two domains by a shift equivalent to counterclockwise rotation of each by 24 degrees. These results suggest that transmembrane signaling by the ANP receptor is initiated via a hormone-induced rotation mechanism.
- Lerner Research Institute, Cleveland Clinic Foundation, Cleveland, OH 44195, USA.
Organizational Affiliation: