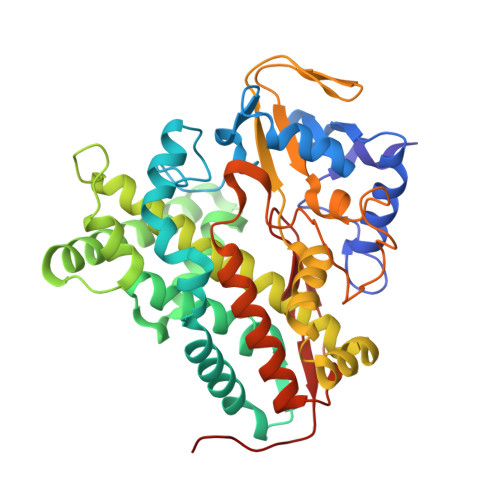
Crystal structure of P450cin in a complex with its substrate, 1,8-cineole, a close structural homologue to D-camphor, the substrate for P450cam
Meharenna, Y.T., Li, H., Hawkes, D.B., Pearson, A.G., De Voss, J., Poulos, T.L.(2004) Biochemistry 43: 9487-9494
- PubMed: 15260491 Search on PubMed
- DOI: https://doi.org/10.1021/bi049293p
- Primary Citation Related Structures:
1T2B - PubMed Abstract:
Cytochrome P450cin catalyzes the monooxygenation of 1,8-cineole, which is structurally very similar to d-camphor, the substrate for the most thoroughly investigated cytochrome P450, cytochrome P450cam. Both 1,8-cineole and d-camphor are C(10) monoterpenes containing a single oxygen atom with very similar molecular volumes. The cytochrome P450cin-substrate complex crystal structure has been solved to 1.7 A resolution and compared with that of cytochrome P450cam. Despite the similarity in substrates, the active site of cytochrome P450cin is substantially different from that of cytochrome P450cam in that the B' helix, essential for substrate binding in many cytochrome P450s including cytochrome P450cam, is replaced by an ordered loop that results in substantial changes in active site topography. In addition, cytochrome P450cin does not have the conserved threonine, Thr252 in cytochrome P450cam, which is generally considered as an integral part of the proton shuttle machinery required for oxygen activation. Instead, the analogous residue in cytochrome P450cin is Asn242, which provides the only direct protein H-bonding interaction with the substrate. Cytochrome P450cin uses a flavodoxin-like redox partner to reduce the heme iron rather than the more traditional ferredoxin-like Fe(2)S(2) redox partner used by cytochrome P450cam and many other bacterial P450s. It thus might be expected that the redox partner docking site of cytochrome P450cin would resemble that of cytochrome P450BM3, which also uses a flavodoxin-like redox partner. Nevertheless, the putative docking site topography more closely resembles cytochrome P450cam than cytochrome P450BM3.
- Department of Molecular Biology, University of California, Irvine, Irvine, California 92697-3900, USA.
Organizational Affiliation: