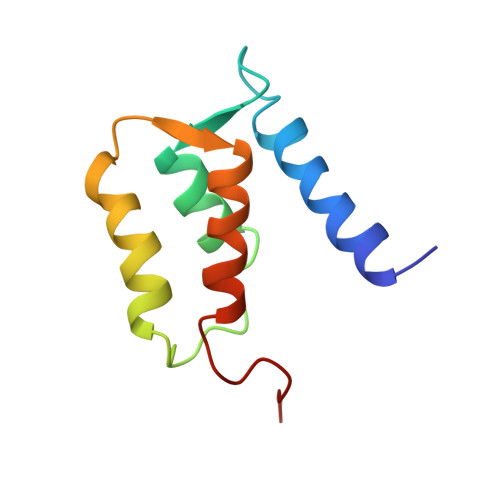
Solution structure of rat apo-S100B(beta beta) as determined by NMR spectroscopy.
Drohat, A.C., Amburgey, J.C., Abildgaard, F., Starich, M.R., Baldisseri, D., Weber, D.J.(1996) Biochemistry 35: 11577-11588
- PubMed: 8794737
- DOI: https://doi.org/10.1021/bi9612226
- Primary Citation of Related Structures:
1SYM - PubMed Abstract:
S100B(beta beta), a member of the S100 protein family, is a Ca(2+)-binding protein with noncovalent interactions at its dimer interface. Each apo-S100 beta subunit (91 residues) has four alpha-helices and a small antiparallel beta-sheet, consistent with two predicted helix-loop-helix Ca(2+)-binding domains known as EF-hands [Amburgey et al. (1995) J. Biomol. NMR 6, 171-179]. The three-dimensional solution structure of apo-S100B(beta beta) from rat has been determined using 2672 distance (14.7 per residue) and 88 dihedral angle restraints derived from multidimensional nuclear magnetic resonance spectroscopy. Apo-S100B (beta beta) is found to be globular and compact with an extensive hydrophobic core and a highly charged surface, consistent with its high solubility. At the symmetric dimer interface, 172 intermolecular nuclear Overhauser effect correlations (NOEs) define the antiparallel alignment of helix I with I' and of helix IV with IV'. The perpendicular association of these pairs of antiparallel helices forms an X-type four-helical bundle at the dimer interface. Whereas, the four helices within each apo-S100 beta subunit adopt a unicornate-type four-helix bundle, with helix I protruding from the parallel bundle of helices II, III, and IV. Accordingly, the orientation of helix III relative to helices I, II, and IV in each subunit differs significantly from that known for other Ca(2+)-binding proteins. Indeed, the interhelical angle (omega) observed in the C-terminal EF-hand of apo-S100 beta is -142 degrees, whereas omega ranges from 118 degrees to 145 degrees in the apo state and from 84 degrees to 128 degrees in the Ca(2+)-bound state for the EF-hands of calbindin D9k, calcyclin, and calmodulin. Thus, a significant conformational change in the C-terminal EF-hand would be required for it to adopt a structure typical of the Ca(2+)-bound state, which could readily explain the dramatic spectral effects observed upon the addition of Ca2+ to apo-S100B(beta beta).
- Department of Biochemistry and Molecular Biology, University of Maryland School of Medicine, Baltimore 21201, USA.
Organizational Affiliation: