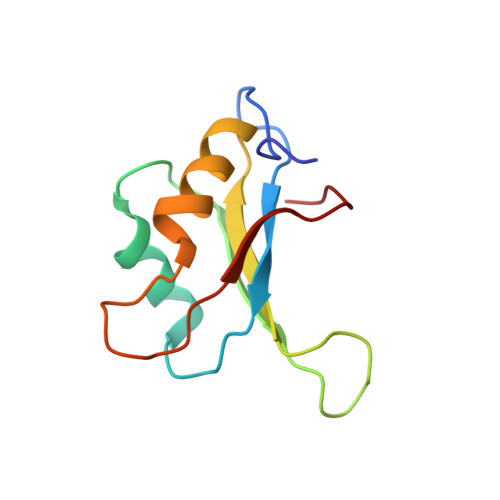
Resonance assignments and solution structure of the second RNA-binding domain of sex-lethal determined by multidimensional heteronuclear magnetic resonance.
Lee, A.L., Kanaar, R., Rio, D.C., Wemmer, D.E.(1994) Biochemistry 33: 13775-13786
- PubMed: 7524663
- DOI: https://doi.org/10.1021/bi00250a031
- Primary Citation Related Structures:
1SXL - PubMed Abstract:
The RNA-binding protein Sex-lethal (Sxl) is a critical regulator of sexual differentiation and dosage compensation in Drosophila. This regulatory activity is a consequence of the ability of Sxl to bind uridine-rich RNA tracts involved in pre-mRNA splicing. Sxl contains two RNP consensus-type RNA-binding domains (RBDs). A structural study of a portion of Sxl (amino acids 199-294) containing the second RNA-binding domain (RBD-2) using multidimensional heteronuclear NMR is presented here. Nearly complete 1H, 13C, and 15N resonance assignments have been obtained from 15N- and 13C/15N-uniformly labeled protein. These assignments were used to analyze 3D 15N-separated NOESY and 13C/13C-separated 4D NOESY spectra which produced 494 total and 169 long-range NOE-derived distance restraints. Along with 41 backbone dihedral restraints, these distance restraints were employed to generate an intermediate-resolution family of calculated structures, which exhibits the beta alpha beta-beta alpha beta tertiary fold found in other RBD-containing proteins. The RMSD to the average structure for the backbone atoms of residues 11-93 is 1.55 +/- 0.30 A, while the RMSD for backbone atoms involved in secondary structure is 0.76 +/- 0.14 A. A capping box [Harper, E.T., & Rose, G.D. (1993) Biochemistry 32, 7605-7609] was identified at the N-terminus of the first helix and has been characterized by short- and medium-range NOEs. Finally, significant structural similarities and differences between Sxl RBD-2 and other RBD-containing proteins are discussed.
- Department of Chemistry, University of California, Berkeley 94720-1460.
Organizational Affiliation: