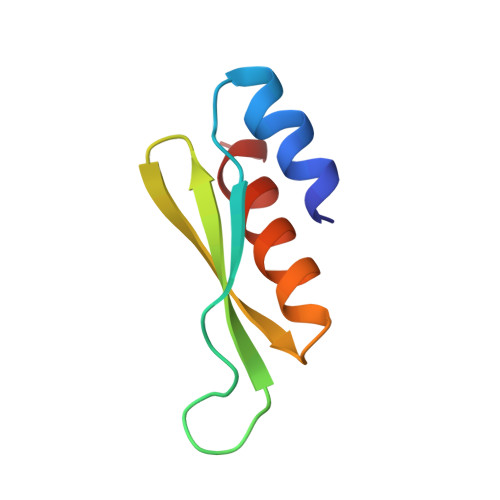
NMR solution structure of a dsRNA binding domain from Drosophila staufen protein reveals homology to the N-terminal domain of ribosomal protein S5.
Bycroft, M., Grunert, S., Murzin, A.G., Proctor, M., St Johnston, D.(1995) EMBO J 14: 3563-3571
- PubMed: 7628456 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/j.1460-2075.1995.tb07362.x
- Primary Citation Related Structures:
1STU - PubMed Abstract:
The double-stranded RNA binding domain (dsRBD) is an approximately 65 amino acid motif that is found in a variety of proteins that interact with double-stranded (ds) RNA, such as Escherichia coli RNase III and the dsRNA-dependent kinase, PKR. Drosophila staufen protein contains five copies of this motif, and the third of these binds dsRNA in vitro. Using multinuclear/multidimensional NMR methods, we have determined that staufen dsRBD3 forms a compact protein domain with an alpha-beta-beta-beta-alpha structure in which the two alpha-helices lie on one face of a three-stranded anti-parallel beta-sheet. This structure is very similar to that of the N-terminal domain of a prokaryotic ribosomal protein S5. Furthermore, the consensus derived from all known S5p family sequences shares several conserved residues with the dsRBD consensus sequence, indicating that the two domains share a common evolutionary origin. Using in vitro mutagenesis, we have identified several surface residues which are important for the RNA binding of the dsRBD, and these all lie on the same side of the domain. Two residues that are essential for RNA binding, F32 and K50, are also conserved in the S5 protein family, suggesting that the two domains interact with RNA in a similar way.
- Cambridge Centre for Protein Engineering, Department of Chemistry, University of Cambridge, UK.
Organizational Affiliation: