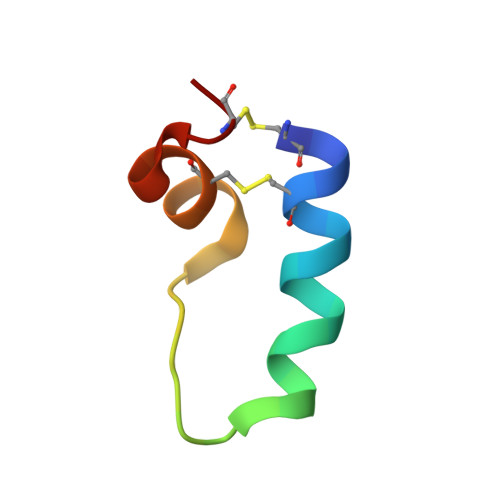
The role of charged amphipathic helices in the structure and function of surfactant protein B.
Waring, A.J., Walther, F.J., Gordon, L.M., Hernandez-Juviel, J.M., Hong, T., Sherman, M.A., Alonso, C., Alig, T., Braun, A., Bacon, D., Zasadzinski, J.A.(2005) J Pept Res 66: 364-374
- PubMed: 16316452 Search on PubMed
- DOI: https://doi.org/10.1111/j.1399-3011.2005.00300.x
- Primary Citation Related Structures:
1SSZ - PubMed Abstract:
Surfactant protein B (SP-B) is essential for normal lung surfactant function. Theoretical models predict that the disulfide cross-linked, N- and C-terminal domains of SP-B fold as charged amphipathic helices, and suggest that these adjacent helices participate in critical surfactant activities. This hypothesis is tested using a disulfide-linked construct (Mini-B) based on the primary sequences of the N- and C-terminal domains. Consistent with theoretical predictions of the full-length protein, both isotope-enhanced Fourier transform infrared (FTIR) spectroscopy and molecular modeling confirm the presence of charged amphipathic alpha-helices in Mini-B. Similar to that observed with native SP-B, Mini-B in model surfactant lipid mixtures exhibits marked in vitro activity, with spread films showing near-zero minimum surface tensions during cycling using captive bubble surfactometry. In vivo, Mini-B shows oxygenation and dynamic compliance that compare favorably with that of full-length SP-B. Mini-B variants (i.e. reduced disulfides or cationic residues replaced by uncharged residues) or Mini-B fragments (i.e. unlinked N- and C-terminal domains) produced greatly attenuated in vivo and in vitro surfactant properties. Hence, the combination of structure and charge for the amphipathic alpha-helical N- and C-terminal domains are key to SP-B function.
- Department of Medicine, Division of Infectious Diseases, UCLA School of Medicine, Center for Health Sciences, Los Angeles, CA 90095, USA. awaring@ucla.edu
Organizational Affiliation: