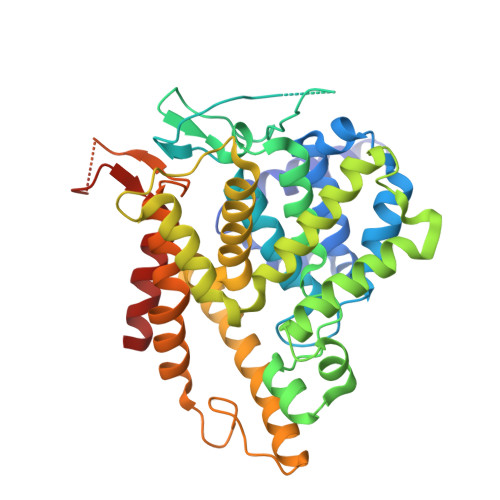
Crystal Structure of Human Phosphodiesterase 3B: Atomic Basis for Substrate and Inhibitor Specificity
Scapin, G., Patel, S.B., Chung, C., Varnerin, J.P., Edmondson, S.D., Mastracchio, A., Parmee, E.R., Singh, S.B., Becker, J.W., Van Der Ploeg, L.H., Tota, M.R.(2004) Biochemistry 43: 6091-6100
- PubMed: 15147193 Search on PubMed
- DOI: https://doi.org/10.1021/bi049868i
- Primary Citation Related Structures:
1SO2, 1SOJ - PubMed Abstract:
Phosphodiesterases (PDEs) are enzymes that modulate cyclic nucleotide signaling and as such are clinical targets for a range of disorders including congestive heart failure, erectile dysfunction, and inflammation. The PDE3 family comprises two highly homologous subtypes expressed in different tissues, and inhibitors of this family have been shown to increase lipolysis in adipocytes. A specific PDE3B (the lipocyte-localized subtype) inhibitor would be a very useful tool to evaluate the effects of PDE3 inhibition on lipolysis and metabolic rate and might become a novel tool for treatment of obesity. We report here the three-dimensional structures of the catalytic domain of human PDE3B in complex with a generic PDE inhibitor and a novel PDE3 selective inhibitor. These structures explain the dual cAMP/cGMP binding capabilities of PDE3, provide the molecular basis for inhibitor specificity, and can supply a valid platform for the design of improved compounds.
- Departments of Medicinal Chemistry, Merck & Co., Rahway, New Jersey 07065, USA. giovanna_scapin@merck.com
Organizational Affiliation: