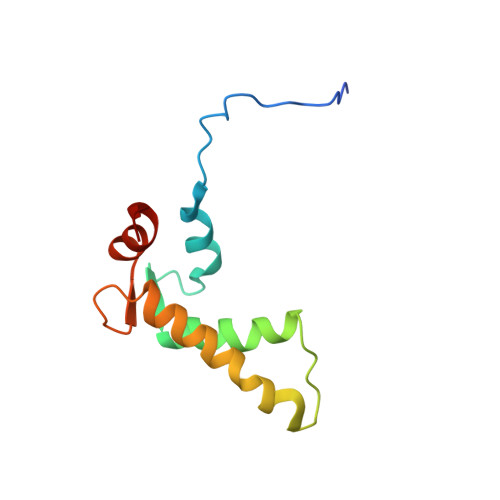
Structural Studies on the Ca(2+)-binding Domain of Human Nucleobindin (Calnuc).
De Alba, E., Tjandra, N.(2004) Biochemistry 43: 10039-10049
- PubMed: 15287731 Search on PubMed
- DOI: https://doi.org/10.1021/bi049310a
- Primary Citation Related Structures:
1SNL - PubMed Abstract:
Nucleobindin, also known as calnuc, participates in Ca2+ storage in the Golgi, as well as in other biological processes that involve DNA-binding and protein-protein interactions. We have determined the three-dimensional solution structure of the Ca(2+)-binding domain of nucleobindin by NMR showing that it consists of two EF-hand motifs. The NMR structure indicates that the phi and psi angles of residues in both motifs are very similar, despite the noncanonical sequence of the C-terminal EF-hand, which contains an arginine residue instead of the typical glycine at the sixth position of the 12-residue loop. The relative orientation of the alpha-helices in the N-terminal EF-hand falls within the common arrangement found in most EF-hand structures. In contrast, the noncanonical EF-hand deviates from the average orientation. The two helix-loop-helix moieties are in the open conformation characteristic of the Ca(2+)-bound state. We find that both motifs bind Ca2+ with apparent dissociation constants of 47 and 40 microM for the noncanonical and the canonical EF-hand, respectively. The Ca(2+)-binding domain of nucleobindin is unstructured in the absence of Ca2+ and folds upon Ca2+ addition. NMR relaxation data and structural studies of the folded domain indicate that it undergoes slow dynamics, suggesting that it is floppier and less compact than a globular domain.
- Laboratory of Biophysical Chemistry, National Heart, Lung, and Blood Institute, National Institutes of Health, 50 Center Drive, Bethesda, Maryland 20892, USA.
Organizational Affiliation: