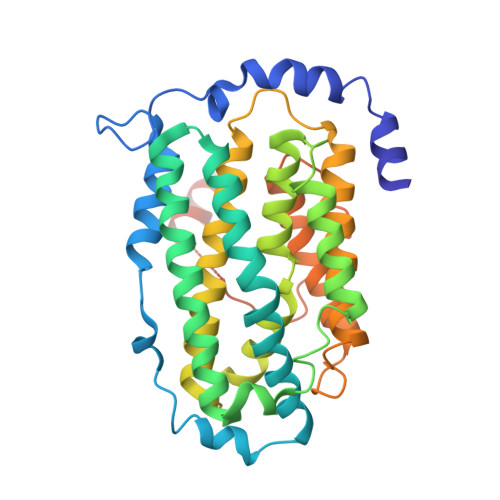
Structures of the yeast ribonucleotide reductase Rnr2 and Rnr4 homodimers.
Sommerhalter, M., Voegtli, W.C., Perlstein, D.L., Ge, J., Stubbe, J., Rosenzweig, A.C.(2004) Biochemistry 43: 7736-7742
- PubMed: 15196016 Search on PubMed
- DOI: https://doi.org/10.1021/bi049510m
- Primary Citation Related Structures:
1SMQ, 1SMS - PubMed Abstract:
Class I ribonucleotide reductases (RNRs) catalyze the reduction of ribonucleotides to deoxyribonucleotides. Eukaryotic RNRs comprise two subunits, the R1 subunit, which contains substrate and allosteric effector binding sites, and the R2 subunit, which houses a catalytically essential diiron-tyrosyl radical cofactor. In Saccharomyces cerevisiae, there are two variants of the R2 subunit, called Rnr2 and Rnr4. Rnr4 is unique in that it lacks three iron-binding residues conserved in all other R2s. Nevertheless, Rnr4 is required to activate Rnr2, and the functional species in vivo is believed to be a heterodimeric complex between the two proteins. The crystal structures of the Rnr2 and Rnr4 homodimers have been determined and are compared to that of the heterodimer. The homodimers are very similar to the heterodimer and to mouse R2 in overall fold, but there are several key differences. In the Rnr2 homodimer, one of the iron-binding helices, helix alphaB, is not well-ordered. In the heterodimer, interactions with a loop region connecting Rnr4 helices alphaA and alpha3 stabilize this Rnr2 helix, which donates iron ligand Asp 145. Sequence differences between Rnr2 and Rnr4 prevent the same interactions from occurring in the Rnr2 homodimer. These findings provide a structural rationale for why the heterodimer is the preferred complex in vivo. The active-site region in the Rnr4 homodimer reveals interactions not apparent in the heterodimer, supporting previous conclusions that this subunit does not bind iron. When taken together, these results support a model in which Rnr4 stabilizes Rnr2 for cofactor assembly and activity.
- Department of Biochemistry, Northwestern University, Evanston, Illinois 60208, USA.
Organizational Affiliation: