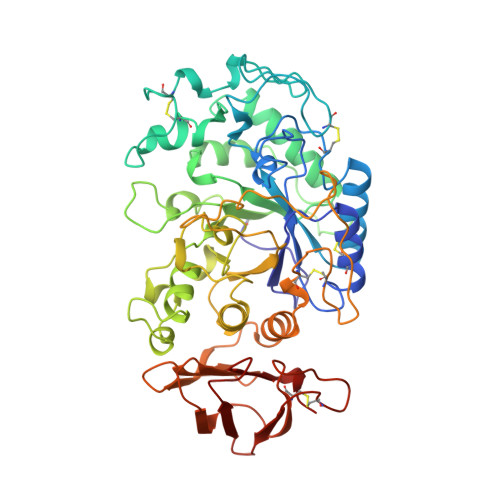
Structure of human salivary alpha-amylase at 1.6 A resolution: implications for its role in the oral cavity.
Ramasubbu, N., Paloth, V., Luo, Y., Brayer, G.D., Levine, M.J.(1996) Acta Crystallogr D Biol Crystallogr 52: 435-446
- PubMed: 15299664 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444995014119
- Primary Citation Related Structures:
1SMD - PubMed Abstract:
Salivary alpha-amylase, a major component of human saliva, plays a role in the initial digestion of starch and may be involved in the colonization of bacteria involved in early dental plaque formation. The three-dimensional atomic structure of salivary amylase has been determined to understand the structure-function relationships of this enzyme. This structure was refined to an R value of 18.4% with 496 amino-acid residues, one calcium ion, one chloride ion and 170 water molecules. Salivary amylase folds into a multidomain structure consisting of three domains, A, B and C. Domain A has a (beta/alpha)(8-) barrel structure, domain B has no definite topology and domain C has a Greek-key barrel structure. The Ca(2+) ion is bound to Asnl00, Arg158, Asp167, His201 and three water molecules. The Cl(-) ion is bound to Arg195, Asn298 and Arg337 and one water molecule. The highly mobile glycine-rich loop 304-310 may act as a gateway for substrate binding and be involved in a 'trap-release' mechanism in the hydrolysis of substrates. Strategic placement of calcium and chloride ions, as well as histidine and tryptophan residues may play a role in differentiating between the glycone and aglycone ends of the polysaccharide substrates. Salivary amylase also possesses a suitable site for binding to enamel surfaces and provides potential sites for the binding of bacterial adhesins.
- Department of Oral Biology and Dental Research Institute, School of Dental Medicine, State University of New York at Buffalo, BY 14214, USA. subbu@crystal1.sdm.buffalo.edu
Organizational Affiliation: