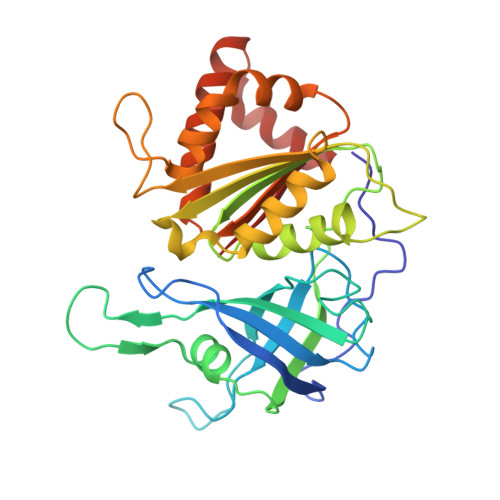
Crystal structure of paprika ferredoxin-NADP+ reductase. Implications for the electron transfer pathway.
Dorowski, A., Hofmann, A., Steegborn, C., Boicu, M., Huber, R.(2001) J Biological Chem 276: 9253-9263
- PubMed: 11053431 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M004576200
- Primary Citation Related Structures:
1SM4 - PubMed Abstract:
cDNA of Capsicum annuum Yolo Wonder (paprika) has been prepared from total cellular RNA, and the complete gene encoding paprika ferredoxin-NADP(+) reductase (pFNR) precursor was sequenced and cloned from this cDNA. Fusion to a T7 promoter allowed expression in Escherichia coli. Both native and recombinant pFNR were purified to homogeneity and crystallized. The crystal structure of pFNR has been solved by Patterson search techniques using the structure of spinach ferredoxin-NADP(+) reductase as search model. The structure was refined at 2.5-A resolution to a crystallographic R-factor of 19.8% (R(free) = 26.5%). The overall structure of pFNR is similar to other members of the ferredoxin-NADP(+) reductase family, the major differences concern a long loop (residues 167-177) that forms part of the FAD binding site and some of the variable loops in surface regions. The different orientation of the FAD binding loop leads to a tighter interaction between pFNR and the adenine moiety of FAD. The physiological redox partners [2Fe-2S]-ferredoxin I and NADP(+) were modeled into the native structure of pFNR. The complexes reveal a protein-protein interaction site that is consistent with existing biochemical data and imply possible orientations for the side chain of tyrosine 362, which has to be displaced by the nicotinamide moiety of NADP(+) upon binding. A reasonable electron transfer pathway could be deduced from the modeled structures of the complexes.
- Max-Planck-Institut für Biochemie, Abteilung Strukturforschung, Am Klopferspitz 18a, 82152 Planegg-Martinsried, Germany. anja.dorowski@gmx.de
Organizational Affiliation: