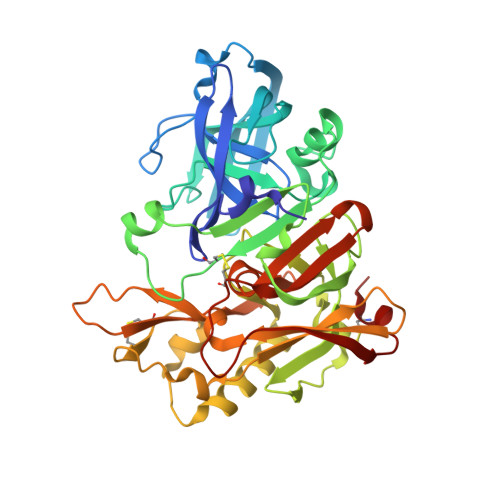
Flap Position of Free Memapsin 2 (beta-Secretase), a Model for Flap Opening in Aspartic Protease Catalysis
Hong, L., Tang, J.(2004) Biochemistry 43: 4689-4695
- PubMed: 15096037 Search on PubMed
- DOI: https://doi.org/10.1021/bi0498252
- Primary Citation Related Structures:
1SGZ - PubMed Abstract:
The three-dimensional structure of unbound human memapsin 2 (beta-secretase) protease domain determined at 2.0-A resolution has revealed a new position of the flap region, which appears to be locked in an "open" position. While the structure outside of the flap is essentially the same as the structure of memapsin 2 bound to an inhibitor, the flap positions are 4.5 A different at the tips. The open position of the flap in the current structure is stabilized by two newly formed intraflap hydrogen bonds and anchored by a new hydrogen bond involving the side chain of Tyr 71 (Tyr 75 in pepsin numbering) in a novel orientation. In molecular modeling experiments, the opening of the flap, 6.5 A at the narrowest point, permits entrance of substrates into the cleft. The narrowest point of the opening may function to discriminate among substrates based on sequence and shape. The observed flap opening may also serve as a model for the flap movement in the catalytic mechanism of eukaryotic aspartic proteases and provide insight for the side-chain selection in the design of memapsin 2 inhibitors.
- Protein Studies Program, Oklahoma Medical Research Foundation, Oklahoma City, Oklahoma 73104, USA.
Organizational Affiliation: