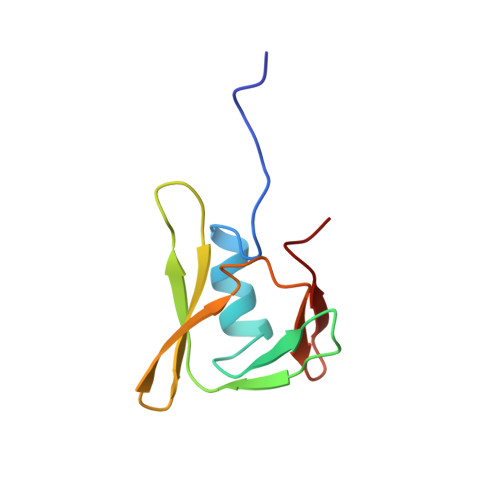
Solution Structure of YaeO, a Rho-specific Inhibitor of Transcription Termination
Gutierrez, P., Kozlov, G., Gabrielli, L., Elias, D., Osborne, M.J., Gallouzi, I.E., Gehring, K.(2007) J Biological Chem 282: 23348-23353
- PubMed: 17565995 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M702010200
- Primary Citation Related Structures:
1SG5 - PubMed Abstract:
Rho-dependent transcription termination is an essential process for the regulation of bacterial gene expression. Thus far, only two Rho-specific inhibitors of bacterial transcription termination have been described, the psu protein from the satellite bacteriophage P4 and YaeO from Escherichia coli. Here, we report the solution structure of YaeO, the first of a Rho-specific inhibitor of transcription termination. YaeO is an acidic protein composed of an N-terminal helix and a seven-stranded beta sandwich. NMR chemical shift perturbation experiments revealed that YaeO binds proximal to the primary nucleic acid binding site of Rho. Based on the NMR titrations, a docked model of the YaeO-Rho complex was calculated. These results suggest that YaeO binds outside the Rho hexamer, acting as a competitive inhibitor of RNA binding. In vitro gel shift assays confirmed the inhibition of nucleic acid binding to Rho. Site-directed mutagenesis showed that the negative character of YaeO is essential for its function in vivo.
- Department of Biochemistry, McGill University, Montreal, Quebec H3G 1Y6, Canada.
Organizational Affiliation: