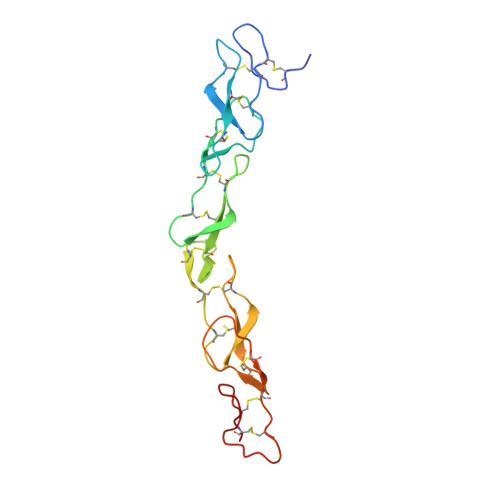
Structure of nerve growth factor complexed with the shared neurotrophin receptor p75
He, X.L., Garcia, K.C.(2004) Science 304: 870-875
- PubMed: 15131306 Search on PubMed
- DOI: https://doi.org/10.1126/science.1095190
- Primary Citation Related Structures:
1SG1 - PubMed Abstract:
Neurotrophins are secreted growth factors critical for the development and maintenance of the vertebrate nervous system. Neurotrophins activate two types of cell surface receptors, the Trk receptor tyrosine kinases and the shared p75 neurotrophin receptor. We have determined the 2.4 A crystal structure of the prototypic neurotrophin, nerve growth factor (NGF), complexed with the extracellular domain of p75. Surprisingly, the complex is composed of an NGF homodimer asymmetrically bound to a single p75. p75 binds along the homodimeric interface of NGF, which disables NGF's symmetry-related second p75 binding site through an allosteric conformational change. Thus, neurotrophin signaling through p75 may occur by disassembly of p75 dimers and assembly of asymmetric 2:1 neurotrophin/p75 complexes, which could potentially engage a Trk receptor to form a trimolecular signaling complex.
- Departments of Microbiology and Immunology, and Structural Biology, Stanford University School of Medicine, Fairchild D319, 299 Campus Drive, Stanford, CA 94305-5124, USA.
Organizational Affiliation: